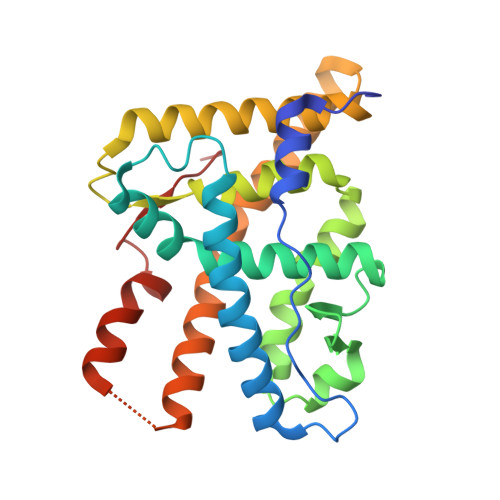
A structural and in vitro characterization of asoprisnil: a selective progesterone receptor modulator.
Madauss, K.P., Grygielko, E.T., Deng, S.J., Sulpizio, A.C., Stanley, T.B., Wu, C., Short, S.A., Thompson, S.K., Stewart, E.L., Laping, N.J., Williams, S.P., Bray, J.D.(2007) Mol Endocrinol 21: 1066-1081
- PubMed: 17356170
- DOI: https://doi.org/10.1210/me.2006-0524
- Primary Citation of Related Structures:
2OVH, 2OVM - PubMed Abstract:
Selective progesterone receptor modulators (SPRMs) have been suggested as therapeutic agents for treatment of gynecological disorders. One such SPRM, asoprisnil, was recently in clinical trials for treatment of uterine fibroids and endometriosis. We present the crystal structures of progesterone receptor (PR) ligand binding domain complexed with asoprisnil and the corepressors nuclear receptor corepressor (NCoR) and SMRT. This is the first report of steroid nuclear receptor crystal structures with ligand and corepressors. These structures show PR in a different conformation than PR complexed with progesterone (P4). We profiled asoprisnil in PR-dependent assays to understand further the PR-mediated mechanism of action. We confirmed previous findings that asoprisnil demonstrated antagonism, but not agonism, in a PR-B transfection assay and the T47D breast cancer cell alkaline phosphatase activity assay. Asoprisnil, but not RU486, weakly recruited the coactivators SRC-1 and AIB1. However, asoprisnil strongly recruited the corepressor NCoR in a manner similar to RU486. Unlike RU486, NCoR binding to asoprisnil-bound PR could be displaced with equal affinity by NCoR or TIF2 peptides. We further showed that it weakly activated T47D cell gene expression of Sgk-1 and PPL and antagonized P4-induced expression of both genes. In rat leiomyoma ELT3 cells, asoprisnil demonstrated partial P4-like inhibition of cyclooxygenase (COX) enzymatic activity and COX-2 gene expression. In the rat uterotrophic assay, asoprisnil demonstrated no P4-like ability to oppose estrogen. Our data suggest that asoprisnil differentially recruits coactivators and corepressors compared to RU486 or P4, and this specific cofactor interaction profile is apparently insufficient to oppose estrogenic activity in rat uterus.
Organizational Affiliation:
Department of Computational, Analytical and Structural Sciences, GlaxoSmithKline Discovery Research, Research Triangle Park, North Carolina 27709, USA.