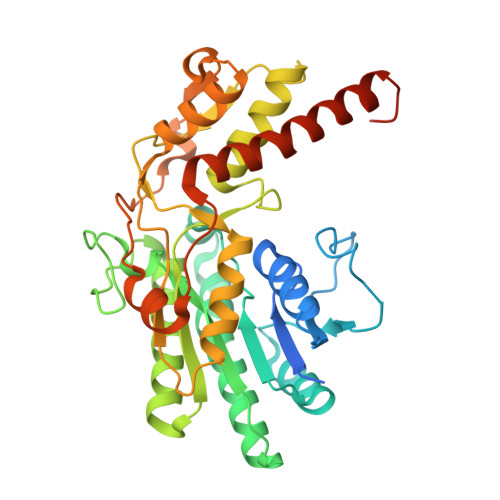
The structure of the MUR1 GDP-mannose 4,6-dehydratase from A. thaliana: Implications for ligand binding and specificity.
Mulichak, A.M., Bonin, C.P., Reiter, W.-D., Garavito, R.M.(2002) Biochemistry 41: 155578-155589
- PubMed: 12501186
- DOI: https://doi.org/10.1021/bi0266683
- Primary Citation of Related Structures:
1N7G, 1N7H - PubMed Abstract:
GDP-D-mannose 4,6-dehydratase catalyzes the first step in the de novo synthesis of GDP-L-fucose, the activated form of L-fucose, which is a component of glycoconjugates in plants known to be important to the development and strength of stem tissues. We have determined the three-dimensional structure of the MUR1 dehydratase isoform from Arabidopsis thaliana complexed with its NADPH cofactor as well as with the ligands GDP and GDP-D-rhamnose. MUR1 is a member of the nucleoside-diphosphosugar modifying subclass of the short-chain dehydrogenase/reductase enzyme family, having homologous structures and a conserved catalytic triad of Lys, Tyr, and Ser/Thr residues. MUR1 is the first member of this subfamily to be observed as a tetramer, the interface of which reveals a close and intimate overlap of neighboring NADP(+)-binding sites. The GDP moiety of the substrate also binds in an unusual syn conformation. The protein-ligand interactions around the hexose moiety of the substrate support the importance of the conserved triad residues and an additional Glu side chain serving as a general base for catalysis. Phe and Arg side chains close to the hexose ring may serve to confer substrate specificity at the O2 position. In the MUR1/GDP-D-rhamnose complex, a single unique monomer within the protein tetramer that has an unoccupied substrate site highlights the conformational changes that accompany substrate binding and may suggest the existence of negative cooperativity in MUR1 function.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, Michigan State University, East Lansing, MI 48824-1319, USA.