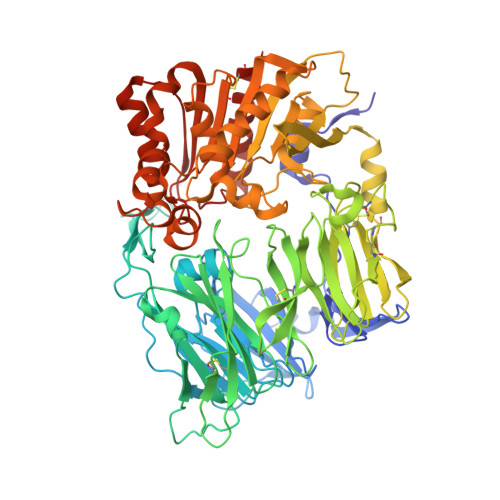
Tyrosine 547 Constitutes an Essential Part of the Catalytic Mechanism of Dipeptidyl Peptidase IV
Bjelke, J.R., Christensen, J., Branner, S., Wagtmann, N., Olsen, C., Kanstrup, A.B., Rasmussen, H.B.(2004) J Biol Chem 279: 34691-34697
- PubMed: 15175333
- DOI: https://doi.org/10.1074/jbc.M405400200
- Primary Citation of Related Structures:
1TK3, 1TKR, 1U8E - PubMed Abstract:
Human dipeptidyl peptidase IV (DPP-IV) is a ubiquitously expressed type II transmembrane serine protease. It cleaves the penultimate positioned prolyl bonds at the N terminus of physiologically important peptides such as the incretin hormones glucagon-like peptide 1 and glucose-dependent insulinotropic peptide. In this study, we have characterized different active site mutants. The Y547F mutant as well as the catalytic triad mutants S630A, D708A, and H740L showed less than 1% wild type activity. X-ray crystal structure analysis of the Y547F mutant revealed no overall changes compared with wild type apoDPP-IV, except the ablation of the hydroxyl group of Tyr(547) and a water molecule positioned in close proximity to Tyr(547). To elucidate further the reaction mechanism, we determined the crystal structure of DPP-IV in complex with diisopropyl fluorophosphate, mimicking the tetrahedral intermediate. The kinetic and structural findings of the tyrosine residue are discussed in relation to the catalytic mechanism of DPP-IV and to the inhibitory mechanism of the 2-cyanopyrrolidine class of potent DPP-IV inhibitors, proposing an explanation for the specificity of this class of inhibitors for the S9b family among serine proteases.
Organizational Affiliation:
Institute of Medical Biochemistry and Genetics, Panum Institute, University of Copenhagen, Bledamsvej 3C, DK-2200 Copenhagen, Denmark.