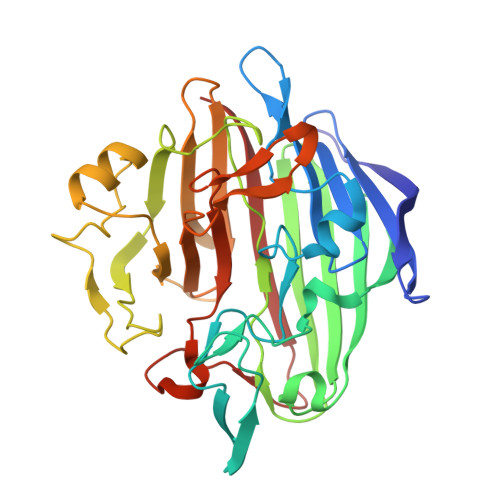
Molecular structure of human galactose mutarotase
Thoden, J.B., Timson, D.J., Reece, R.J., Holden, H.M.(2004) J Biol Chem 279: 23431-23437
- PubMed: 15026423
- DOI: https://doi.org/10.1074/jbc.M402347200
- Primary Citation of Related Structures:
1SNZ, 1SO0 - PubMed Abstract:
Galactose mutarotase catalyzes the conversion of beta-d-galactose to alpha-d-galactose during normal galactose metabolism. The enzyme has been isolated from bacteria, plants, and animals and is present in the cytoplasm of most cells. Here we report the x-ray crystallographic analysis of human galactose mutarotase both in the apoform and complexed with its substrate, beta-d-galactose. The polypeptide chain folds into an intricate array of 29 beta-strands, 25 classical reverse turns, and 2 small alpha-helices. There are two cis-peptide bonds at Arg-78 and Pro-103. The sugar ligand sits in a shallow cleft and is surrounded by Asn-81, Arg-82, His-107, His-176, Asp-243, Gln-279, and Glu-307. Both the side chains of Glu-307 and His-176 are in the proper location to act as a catalytic base and a catalytic acid, respectively. These residues are absolutely conserved among galactose mutarotases. To date, x-ray models for three mutarotases have now been reported, namely that described here and those from Lactococcus lactis and Caenorhabditis elegans. The molecular architectures of these enzymes differ primarily in the loop regions connecting the first two beta-strands. In the human protein, there are six extra residues in the loop compared with the bacterial protein for an approximate longer length of 9 A. In the C. elegans protein, the first 17 residues are missing, thereby reducing the total number of beta-strands by one.
Organizational Affiliation:
Department of Biochemistry, University of Wisconsin, Madison, Wisconsin 53706, USA.