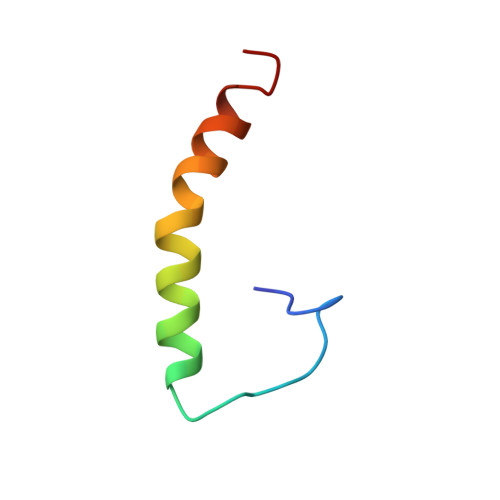
Solution structure of amyloid beta-peptide(1-40) in a water-micelle environment. Is the membrane-spanning domain where we think it is?
Coles, M., Bicknell, W., Watson, A.A., Fairlie, D.P., Craik, D.J.(1998) Biochemistry 37: 11064-11077
- PubMed: 9693002
- DOI: https://doi.org/10.1021/bi972979f
- Primary Citation of Related Structures:
1BA4 - PubMed Abstract:
The three-dimensional solution structure of the 40 residue amyloid beta-peptide, Abeta(1-40), has been determined using NMR spectroscopy at pH 5.1, in aqueous sodium dodecyl sulfate (SDS) micelles. In this environment, which simulates to some extent a water-membrane medium, the peptide is unstructured between residues 1 and 14 which are mainly polar and likely solvated by water. However, the rest of the protein adopts an alpha-helical conformation between residues 15 and 36 with a kink or hinge at 25-27. This largely hydrophobic region is likely solvated by SDS. Based on the derived structures, evidence is provided in support of a possible new location for the transmembrane domain of Abeta within the amyloid precursor protein (APP). Studies between pH 4.2 and 7.9 reveal a pH-dependent helix-coil conformational switch. At the lower pH values, where the carboxylate residues are protonated, the helix is uncharged, intact, and lipid-soluble. As the pH increases above 6. 0, part of the helical region (15-24) becomes less structured, particularly near residues E22 and D23 where deprotonation appears to facilitate unwinding of the helix. This pH-dependent unfolding to a random coil conformation precedes any tendency of this peptide to aggregate to a beta-sheet as the pH increases. The structural biology described herein for Abeta(1-40) suggests that (i) the C-terminal two-thirds of the peptide is an alpha-helix in membrane-like environments, (ii) deprotonation of two acidic amino acids in the helix promotes a helix-coil conformational transition that precedes aggregation, (iii) a mobile hinge exists in the helical region of Abeta(1-40) and this may be relevant to its membrane-inserting properties and conformational rearrangements, and (iv) the location of the transmembrane domain of amyloid precursor proteins may be different from that accepted in the literature. These results may provide new insight to the structural properties of amyloid beta-peptides of relevance to Alzheimer's disease.
Organizational Affiliation:
Centre for Drug Design and Development, The University of Queensland, Brisbane, Australia.