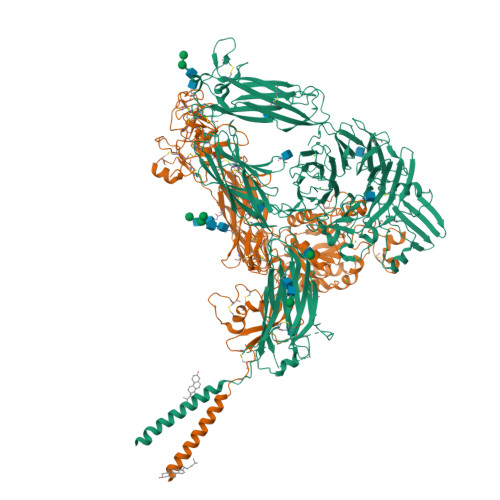
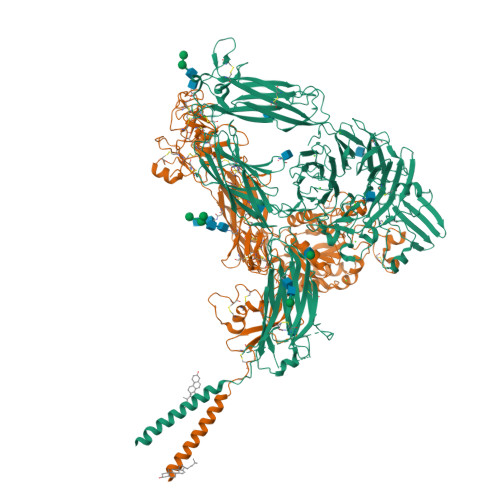
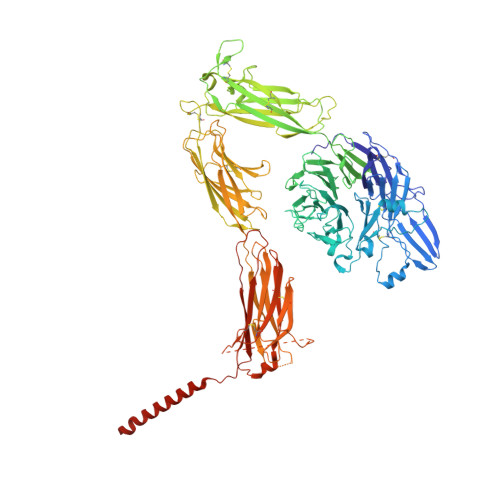
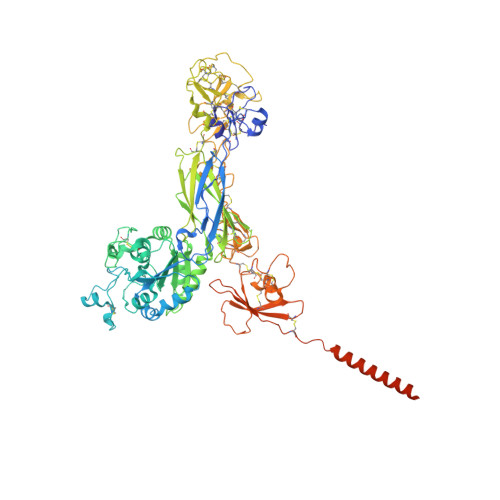
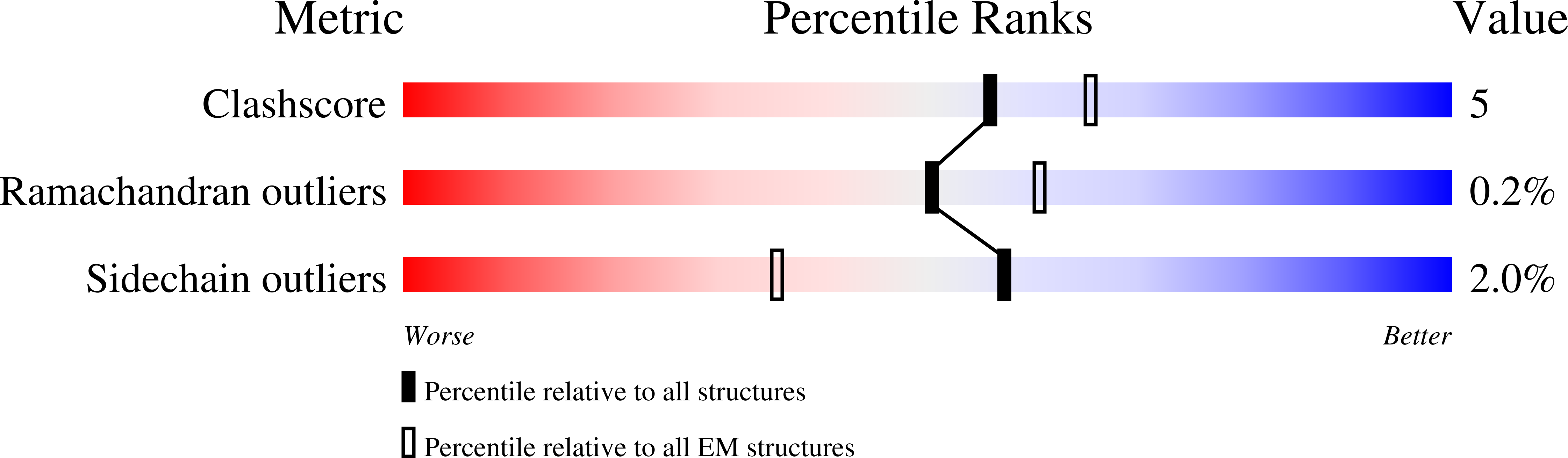Cryo-EM structures of full-length integrin alpha IIb beta 3 in native lipids.
Adair, B.D., Xiong, J.P., Yeager, M., Arnaout, M.A.(2023) Nat Commun 14: 4168-4168
- PubMed: 37443315
- DOI: https://doi.org/10.1038/s41467-023-39763-0
- Primary Citation of Related Structures:
8T2U, 8T2V - PubMed Abstract:
Platelet integrin αIIbβ3 is maintained in a bent inactive state (low affinity to physiologic ligand), but can rapidly switch to a ligand-competent (high-affinity) state in response to intracellular signals ("inside-out" activation). Once bound, ligands drive proadhesive "outside-in" signaling. Anti-αIIbβ3 drugs like eptifibatide can engage the inactive integrin directly, inhibiting thrombosis but inadvertently impairing αIIbβ3 hemostatic functions. Bidirectional αIIbβ3 signaling is mediated by reorganization of the associated αIIb and β3 transmembrane α-helices, but the underlying changes remain poorly defined absent the structure of the full-length receptor. We now report the cryo-EM structures of full-length αIIbβ3 in its apo and eptifibatide-bound states in native cell-membrane nanoparticles at near-atomic resolution. The apo form adopts the bent inactive state but with separated transmembrane α-helices, and a fully accessible ligand-binding site that challenges the model that this site is occluded by the plasma membrane. Bound eptifibatide triggers dramatic conformational changes that may account for impaired hemostasis. These results advance our understanding of integrin structure and function and may guide development of safer inhibitors.
Organizational Affiliation:
Leukocyte Biology and Inflammation Laboratory, Structural Biology Program, Division of Nephrology, Department of Medicine, Massachusetts General Hospital, Boston, Massachusetts, 02114, USA.