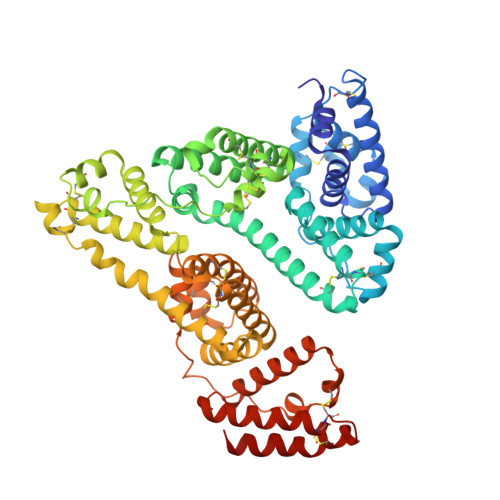
Structural Analysis of Human Serum Albumin in Complex with the Fibrate Drug Gemfibrozil.
Liberi, S., Linciano, S., Moro, G., De Toni, L., Cendron, L., Angelini, A.(2022) Int J Mol Sci 23
- PubMed: 35163693
- DOI: https://doi.org/10.3390/ijms23031769
- Primary Citation of Related Structures:
7QFE - PubMed Abstract:
Gemfibrozil (GEM) is an orally administered lipid-regulating fibrate derivative drug sold under the brand name Lopid ® , among others. Since its approval in the early 80s, GEM has been largely applied to treat hypertriglyceridemia and other disorders of lipid metabolism. Though generally well tolerated, GEM can alter the distribution and the free, active concentration of some co-administered drugs, leading to adverse effects. Most of them appear to be related to the ability of GEM to bind with high affinity human serum albumin (HSA), the major drug-carrier protein in blood plasma. Here, we report the crystal structure of HSA in complex with GEM. Two binding sites have been identified, namely Sudlow's binding sites I (FA7) and II (FA3-FA4). A comparison of the crystal structure of HSA in complex with GEM with those of other previously described HSA-drug complexes enabled us to appreciate the analogies and differences in their respective binding modes. The elucidation of the molecular interaction between GEM and HSA might offer the basis for the development of novel GEM derivatives that can be safely and synergistically co-administered with other drugs, enabling augmented therapeutic efficacies.
Organizational Affiliation:
Department of Biology, University of Padua, Viale G. Colombo 3, 35131 Padua, Italy.