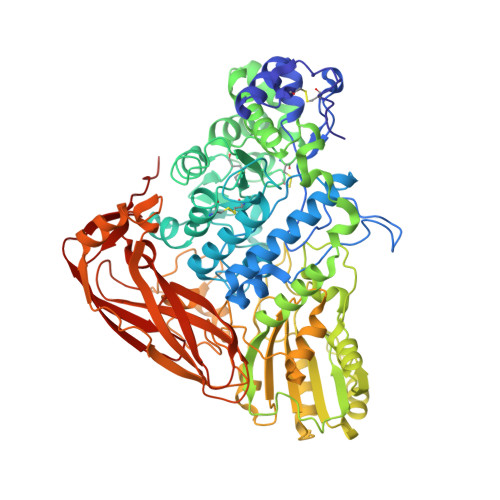
Comparison of glycoside hydrolase family 3 beta-xylosidases from basidiomycetes and ascomycetes reveals evolutionarily distinct xylan degradation systems.
Kojima, K., Sunagawa, N., Mikkelsen, N.E., Hansson, H., Karkehabadi, S., Samejima, M., Sandgren, M., Igarashi, K.(2022) J Biol Chem 298: 101670-101670
- PubMed: 35120929
- DOI: https://doi.org/10.1016/j.jbc.2022.101670
- Primary Citation of Related Structures:
7VC6, 7VC7 - PubMed Abstract:
Xylan is the most common hemicellulose in plant cell walls, though the structure of xylan polymers differs between plant species. Here, to gain a better understanding of fungal xylan degradation systems, which can enhance enzymatic saccharification of plant cell walls in industrial processes, we conducted a comparative study of two glycoside hydrolase family 3 (GH3) β-xylosidases (Bxls), one from the basidiomycete Phanerochaete chrysosporium (PcBxl3), and the other from the ascomycete Trichoderma reesei (TrXyl3A). A comparison of the crystal structures of the two enzymes, both with saccharide bound at the catalytic center, provided insight into the basis of substrate binding at each subsite. PcBxl3 has a substrate-binding pocket at subsite -1, while TrXyl3A has an extra loop that contains additional binding subsites. Furthermore, kinetic experiments revealed that PcBxl3 degraded xylooligosaccharides faster than TrXyl3A, while the K M values of TrXyl3A were lower than those of PcBxl3. The relationship between substrate specificity and degree of polymerization of substrates suggested that PcBxl3 preferentially degrades xylobiose (X 2 ), while TrXyl3A degrades longer xylooligosaccharides. Moreover, docking simulation supported the existence of extended positive subsites of TrXyl3A in the extra loop located at the N-terminus of the protein. Finally, phylogenetic analysis suggests that wood-decaying basidiomycetes use Bxls such as PcBxl3 that act efficiently on xylan structures from woody plants, whereas molds use instead Bxls that efficiently degrade xylan from grass. Our results provide added insights into fungal efficient xylan degradation systems.
Organizational Affiliation:
Department of Biomaterial Sciences, Graduate School of Agricultural and Life Sciences, The University of Tokyo, Bunkyo-ku, Tokyo, Japan.