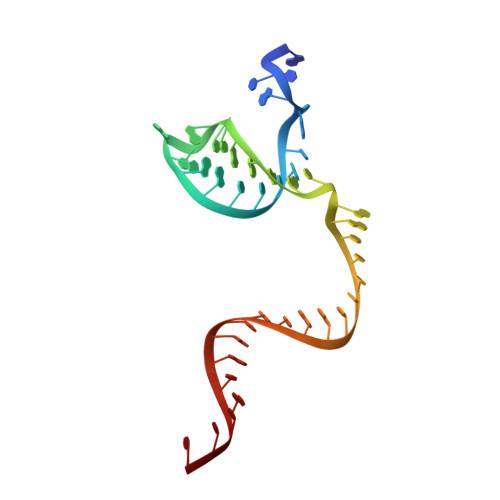
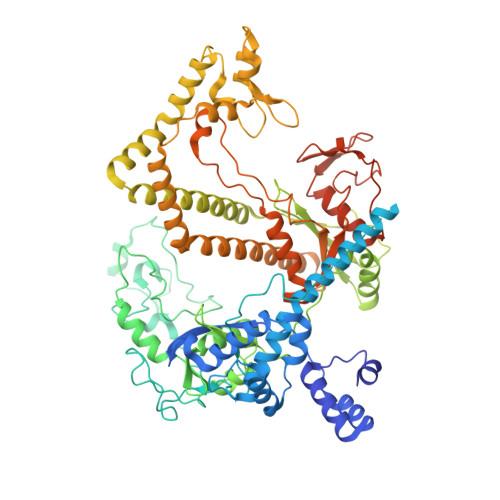
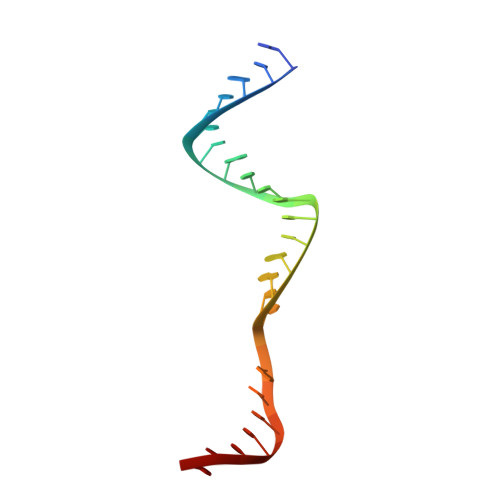
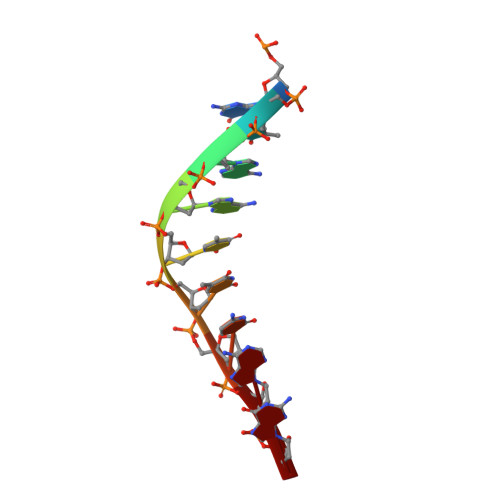
Structure of the mini-RNA-guided endonuclease CRISPR-Cas12j3.
Carabias, A., Fuglsang, A., Temperini, P., Pape, T., Sofos, N., Stella, S., Erlendsson, S., Montoya, G.(2021) Nat Commun 12: 4476-4476
- PubMed: 34294706
- DOI: https://doi.org/10.1038/s41467-021-24707-3
- Primary Citation of Related Structures:
7ODF - PubMed Abstract:
CRISPR-Cas12j is a recently identified family of miniaturized RNA-guided endonucleases from phages. These ribonucleoproteins provide a compact scaffold gathering all key activities of a genome editing tool. We provide the first structural insight into the Cas12j family by determining the cryoEM structure of Cas12j3/R-loop complex after DNA cleavage. The structure reveals the machinery for PAM recognition, hybrid assembly and DNA cleavage. The crRNA-DNA hybrid is directed to the stop domain that splits the hybrid, guiding the T-strand towards the catalytic site. The conserved RuvC insertion is anchored in the stop domain and interacts along the phosphate backbone of the crRNA in the hybrid. The assembly of a hybrid longer than 12-nt activates catalysis through key functional residues in the RuvC insertion. Our findings suggest why Cas12j unleashes unspecific ssDNA degradation after activation. A site-directed mutagenesis analysis supports the DNA cutting mechanism, providing new avenues to redesign CRISPR-Cas12j nucleases for genome editing.
Organizational Affiliation:
Structural Molecular Biology Group, Novo Nordisk Foundation Centre for Protein Research, Faculty of Health and Medical Sciences University of Copenhagen, Copenhagen, Denmark.