Enzymatic Preparation of 2'-5',3'-5'-Cyclic Dinucleotides, Their Binding Properties to Stimulator of Interferon Genes Adaptor Protein, and Structure/Activity Correlations.
Novotna, B., Vanekova, L., Zavrel, M., Budesinsky, M., Dejmek, M., Smola, M., Gutten, O., Tehrani, Z.A., Pimkova Polidarova, M., Brazdova, A., Liboska, R., Stepanek, I., Vavrina, Z., Jandusik, T., Nencka, R., Rulisek, L., Boura, E., Brynda, J., Pav, O., Birkus, G.(2019) J Med Chem 62: 10676-10690
- PubMed: 31715099
- DOI: https://doi.org/10.1021/acs.jmedchem.9b01062
- Primary Citation of Related Structures:
6S26, 6S27 - PubMed Abstract:
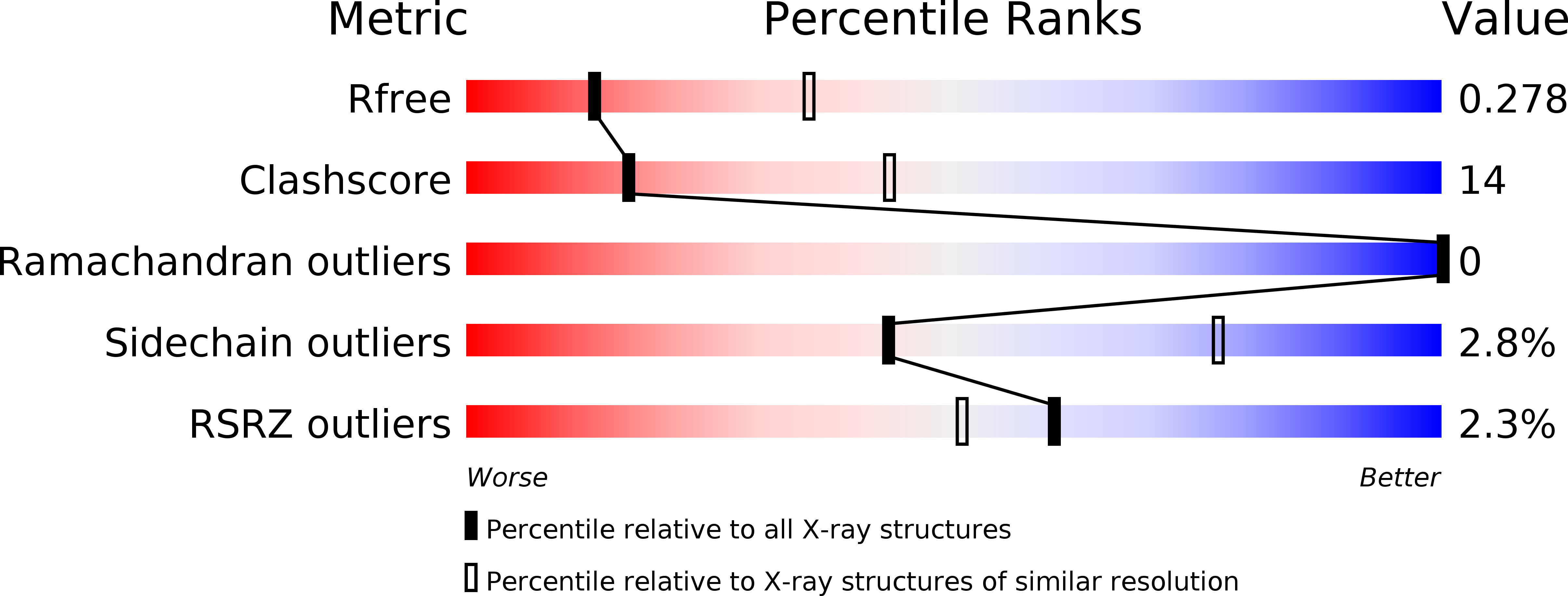
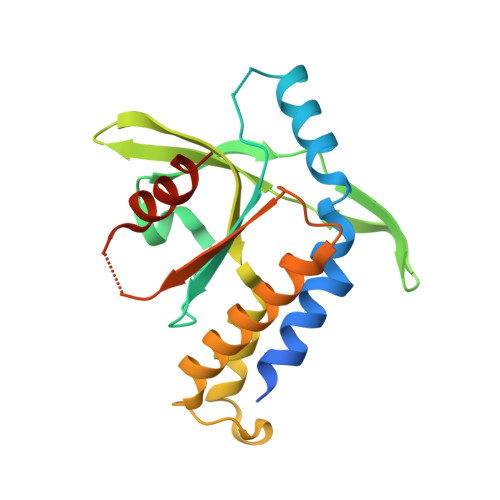
Cyclic dinucleotides are second messengers in the cyclic GMP-AMP synthase (cGAS)-stimulator of interferon genes (STING) pathway, which plays an important role in recognizing tumor cells and viral or bacterial infections. They bind to the STING adaptor protein and trigger expression of cytokines via TANK binding kinase 1 (TBK1)/interferon regulatory factor 3 (IRF3) and inhibitor of nuclear factor-κB (IκB) kinase (IKK)/nuclear factor-κB (NFκB) signaling cascades. In this work, we describe an enzymatic preparation of 2'-5',3'-5'-cyclic dinucleotides (2'3'CDNs) with use of cyclic GMP-AMP synthases (cGAS) from human, mouse, and chicken. We profile substrate specificity of these enzymes by employing a small library of nucleotide-5'-triphosphate (NTP) analogues and use them to prepare 33 2'3'CDNs. We also determine affinity of these CDNs to five different STING haplotypes in cell-based and biochemical assays and describe properties needed for their optimal activity toward all STING haplotypes. Next, we study their effect on cytokine and chemokine induction by human peripheral blood mononuclear cells (PBMCs) and evaluate their cytotoxic effect on monocytes. Additionally, we report X-ray crystal structures of two new CDNs bound to STING protein and discuss structure-activity relationship by using quantum and molecular mechanical (QM/MM) computational modeling.
Organizational Affiliation:
Institute of Organic Chemistry and Biochemistry of the Czech Academy of Sciences and Gilead Sciences Research Centre at IOCB , Flemingovo nam. 2 , Prague 16610 , Czech Republic.