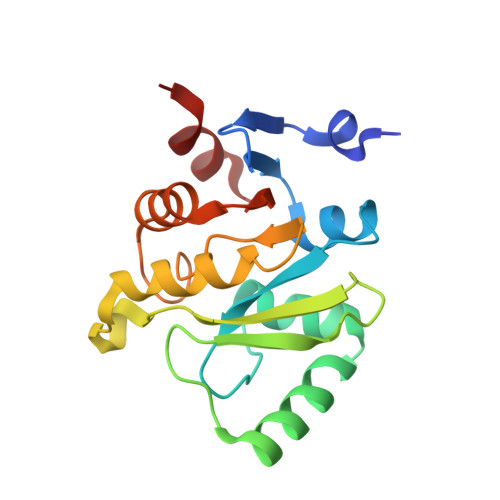
Nsp3 of coronaviruses: Structures and functions of a large multi-domain protein.
Lei, J., Kusov, Y., Hilgenfeld, R.(2018) Antiviral Res 149: 58-74
- PubMed: 29128390
- DOI: https://doi.org/10.1016/j.antiviral.2017.11.001
- Primary Citation of Related Structures:
5HOL - PubMed Abstract:
The multi-domain non-structural protein 3 (Nsp3) is the largest protein encoded by the coronavirus (CoV) genome, with an average molecular mass of about 200 kD. Nsp3 is an essential component of the replication/transcription complex. It comprises various domains, the organization of which differs between CoV genera, due to duplication or absence of some domains. However, eight domains of Nsp3 exist in all known CoVs: the ubiquitin-like domain 1 (Ubl1), the Glu-rich acidic domain (also called "hypervariable region"), a macrodomain (also named "X domain"), the ubiquitin-like domain 2 (Ubl2), the papain-like protease 2 (PL2 pro ), the Nsp3 ectodomain (3Ecto, also called "zinc-finger domain"), as well as the domains Y1 and CoV-Y of unknown functions. In addition, the two transmembrane regions, TM1 and TM2, exist in all CoVs. The three-dimensional structures of domains in the N-terminal two thirds of Nsp3 have been investigated by X-ray crystallography and/or nuclear magnetic resonance (NMR) spectroscopy since the outbreaks of Severe Acute Respiratory Syndrome coronavirus (SARS-CoV) in 2003 as well as Middle-East Respiratory Syndrome coronavirus (MERS-CoV) in 2012. In this review, the structures and functions of these domains of Nsp3 are discussed in depth.
Organizational Affiliation:
Institute of Biochemistry, Center for Structural and Cell Biology in Medicine, University of Lübeck, Ratzeburger Allee 160, 23562 Lübeck, Germany.