Crystal structure of complex
Wang, J., Yuan, Z., Cui, Y., Xie, R., Wang, M., Ma, Y., Yu, X., Liu, X.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Tudor-interacting repair regulator protein | 210 | Homo sapiens | Mutation(s): 0 Gene Names: NUDT16L1, SDOS, TIRR | 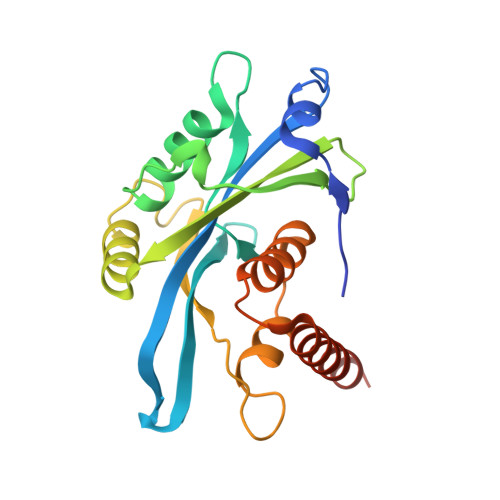 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q9BRJ7 (Homo sapiens) Explore Q9BRJ7 Go to UniProtKB: Q9BRJ7 | |||||
GTEx: ENSG00000168101 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9BRJ7 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| TP53-binding protein 1 | 180 | Homo sapiens | Mutation(s): 0 Gene Names: TP53BP1 | 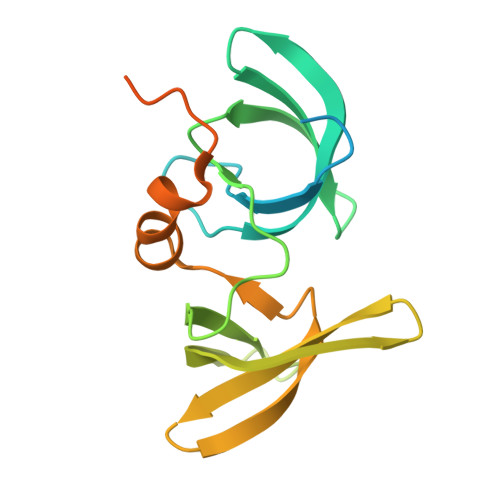 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q12888 (Homo sapiens) Explore Q12888 Go to UniProtKB: Q12888 | |||||
PHAROS: Q12888 GTEx: ENSG00000067369 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q12888 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 111.144 | α = 90 |
| b = 103.922 | β = 95.46 |
| c = 61.47 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-3000 | data reduction |
| HKL-3000 | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Natural Science Foundation of China | China | 81672794 |