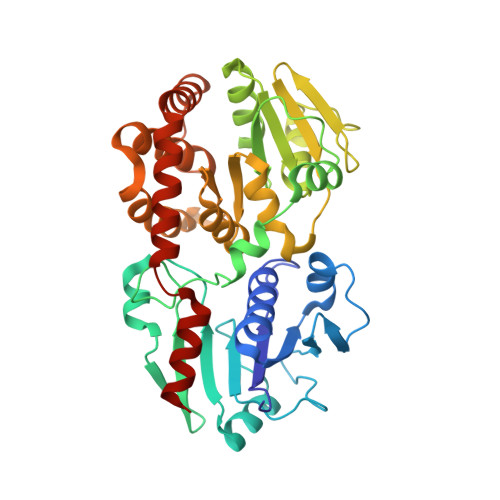
Structural basis of protein arginine rhamnosylation by glycosyltransferase EarP
Sengoku, T., Suzuki, T., Dohmae, N., Watanabe, C., Honma, T., Hikida, Y., Yamaguchi, Y., Takahashi, H., Yokoyama, S., Yanagisawa, T.(2018) Nat Chem Biol 14: 368-374
- PubMed: 29440735
- DOI: https://doi.org/10.1038/s41589-018-0002-y
- Primary Citation of Related Structures:
5WXI, 5WXJ, 5WXK, 5XVR - PubMed Abstract:
Protein glycosylation regulates many cellular processes. Numerous glycosyltransferases with broad substrate specificities have been structurally characterized. A novel inverting glycosyltransferase, EarP, specifically transfers rhamnose from dTDP-β-L-rhamnose to Arg32 of bacterial translation elongation factor P (EF-P) to activate its function. Here we report a crystallographic study of Neisseria meningitidis EarP. The EarP structure contains two tandem Rossmann-fold domains, which classifies EarP in glycosyltransferase superfamily B. In contrast to other structurally characterized protein glycosyltransferases, EarP binds the entire β-sheet structure of EF-P domain I through numerous interactions that specifically recognize its conserved residues. Thus Arg32 is properly located at the active site, and causes structural change in a conserved dTDP-β-L-rhamnose-binding loop of EarP. Rhamnosylation by EarP should occur via an S N 2 reaction, with Asp20 as the general base. The Arg32 binding and accompanying structural change of EarP may induce a change in the rhamnose-ring conformation suitable for the reaction.
Organizational Affiliation:
RIKEN Structural Biology Laboratory, Yokohama, Japan.