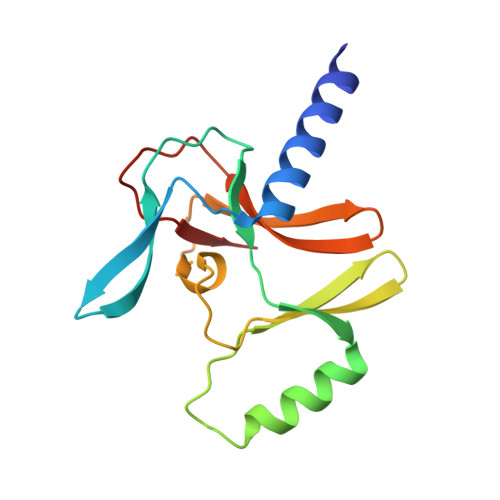
Structure-Based Design of a Covalent Inhibitor of the SET Domain-Containing Protein 8 (SETD8) Lysine Methyltransferase.
Butler, K.V., Ma, A., Yu, W., Li, F., Tempel, W., Babault, N., Pittella-Silva, F., Shao, J., Wang, J., Luo, M., Vedadi, M., Brown, P.J., Arrowsmith, C.H., Jin, J.(2016) J Med Chem 59: 9881-9889
- PubMed: 27804297
- DOI: https://doi.org/10.1021/acs.jmedchem.6b01244
- Primary Citation of Related Structures:
5T5G, 5TH7 - PubMed Abstract:
Selective inhibitors of protein lysine methyltransferases, including SET domain-containing protein 8 (SETD8), are highly desired, as only a fraction of these enzymes are associated with high-quality inhibitors. From our previously discovered SETD8 inhibitor, we developed a more potent analog and solved a cocrystal structure, which is the first crystal structure of SETD8 in complex with a small-molecule inhibitor. This cocrystal structure allowed the design of a covalent inhibitor of SETD8 (MS453), which specifically modifies a cysteine residue near the inhibitor binding site, has an IC 50 value of 804 nM, reacts with SETD8 with near-quantitative yield, and is selective for SETD8 against 28 other methyltransferases. We also solved the crystal structure of the covalent inhibitor in complex with SETD8. This work provides atomic-level perspective on the inhibition of SETD8 by small molecules and will help identify high-quality chemical probes of SETD8.
Organizational Affiliation:
Department of Pharmacological Sciences and Department of Oncological Sciences, Icahn School of Medicine at Mount Sinai , New York, New York 10029, United States.