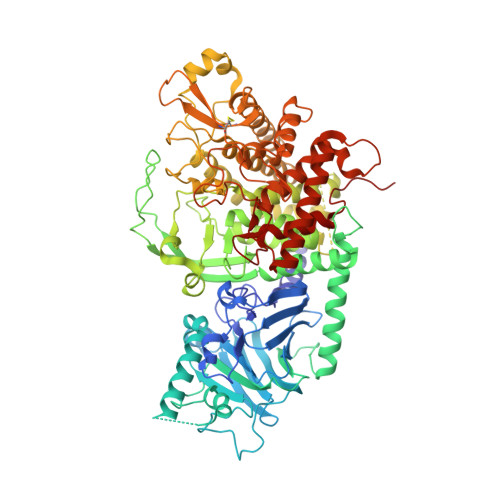
Specificity of processing alpha-glucosidase I is guided by the substrate conformation: crystallographic and in silico studies.
Barker, M.K., Rose, D.R.(2013) J Biol Chem 288: 13563-13574
- PubMed: 23536181
- DOI: https://doi.org/10.1074/jbc.M113.460436
- Primary Citation of Related Structures:
4J5T - PubMed Abstract:
The enzyme “GluI” is key to the synthesis of critical glycoproteins in the cell. We have determined the structure of GluI, and modeled binding with its unique sugar substrate. The specificity of this interaction derives from a unique conformation of the substrate. Understanding the mechanism of the enzyme is of basic importance and relevant to potential development of antiviral inhibitors. Processing α-glucosidase I (GluI) is a key member of the eukaryotic N-glycosylation processing pathway, selectively catalyzing the first glycoprotein trimming step in the endoplasmic reticulum. Inhibition of GluI activity impacts the infectivity of enveloped viruses; however, despite interest in this protein from a structural, enzymatic, and therapeutic standpoint, little is known about its structure and enzymatic mechanism in catalysis of the unique glycan substrate Glc3Man9GlcNAc2. The first structural model of eukaryotic GluI is here presented at 2-Å resolution. Two catalytic residues are proposed, mutations of which result in catalytically inactive, properly folded protein. Using Autodocking methods with the known substrate and inhibitors as ligands, including a novel inhibitor characterized in this work, the active site of GluI was mapped. From these results, a model of substrate binding has been formulated, which is most likely conserved in mammalian GluI.
Organizational Affiliation:
Department of Medical Biophysics, University of Toronto, Ontario Cancer Institute, Princess Margaret Hospital, Toronto, Ontario M5G 2M9, Canada. megan.barker@utoronto.ca