Acinetobacter Baumannii Fold Ligand Complexes; Potent Inhibitors of Folate Metabolism and a Re-Evaluation of the Ly374571 Structure.
Eadsforth, T.C., Maluf, F.V., Hunter, W.N.(2012) FEBS J 279: 4350
- PubMed: 23050773
- DOI: https://doi.org/10.1111/febs.12025
- Primary Citation of Related Structures:
4B4U, 4B4V, 4B4W - PubMed Abstract:
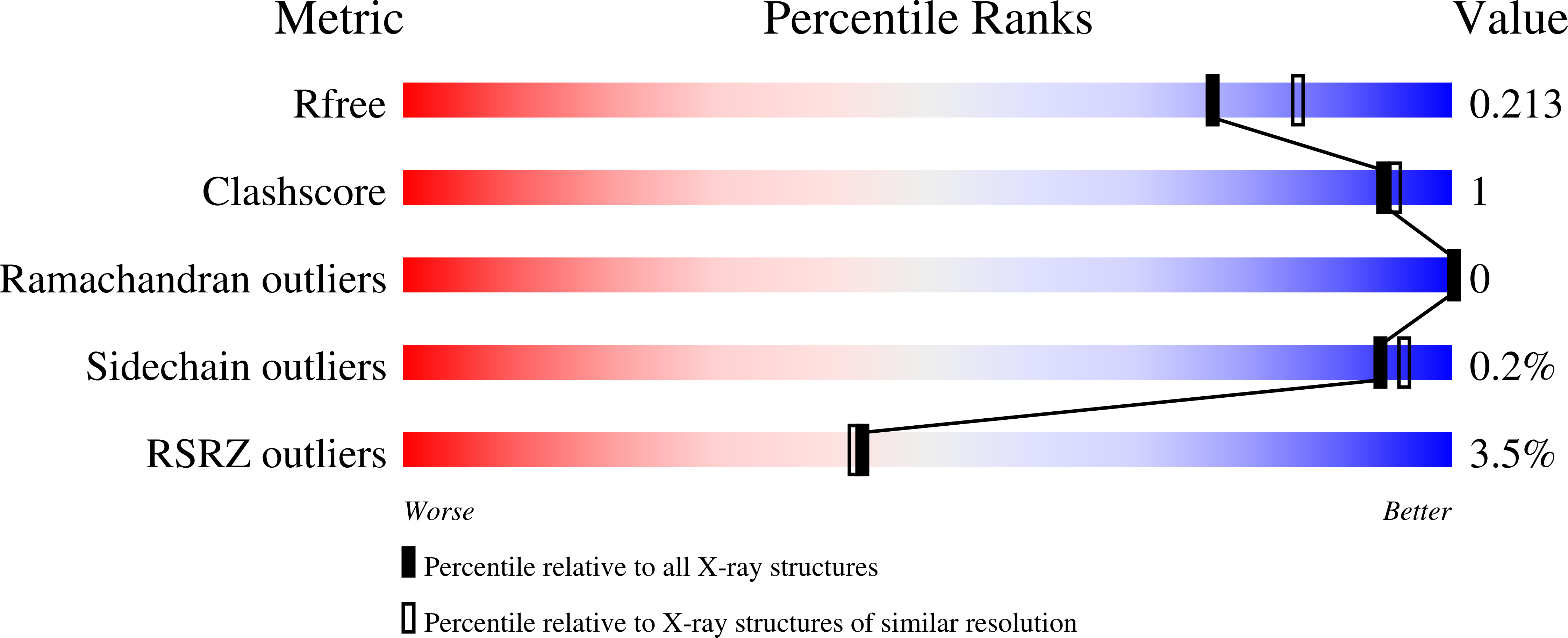
The bifunctional N(5),N(10)-methylenetetrahydrofolate dehydrogenase/cyclohydrolase (DHCH or FolD), which is widely distributed in prokaryotes and eukaryotes, is involved in the biosynthesis of folate cofactors that are essential for growth and cellular development. The enzyme activities represent a potential antimicrobial drug target. We have characterized the kinetic properties of FolD from the Gram-negative pathogen Acinetobacter baumanni and determined high-resolution crystal structures of complexes with a cofactor and two potent inhibitors. The data reveal new details with respect to the molecular basis of catalysis and potent inhibition. A unexpected finding was that our crystallographic data revealed a different structure for LY374571 (an inhibitor studied as an antifolate) than that previously published. The implications of this observation are discussed.
Organizational Affiliation:
Division of Biological Chemistry and Drug Discovery, College of Life Sciences, University of Dundee, Dundee, UK.