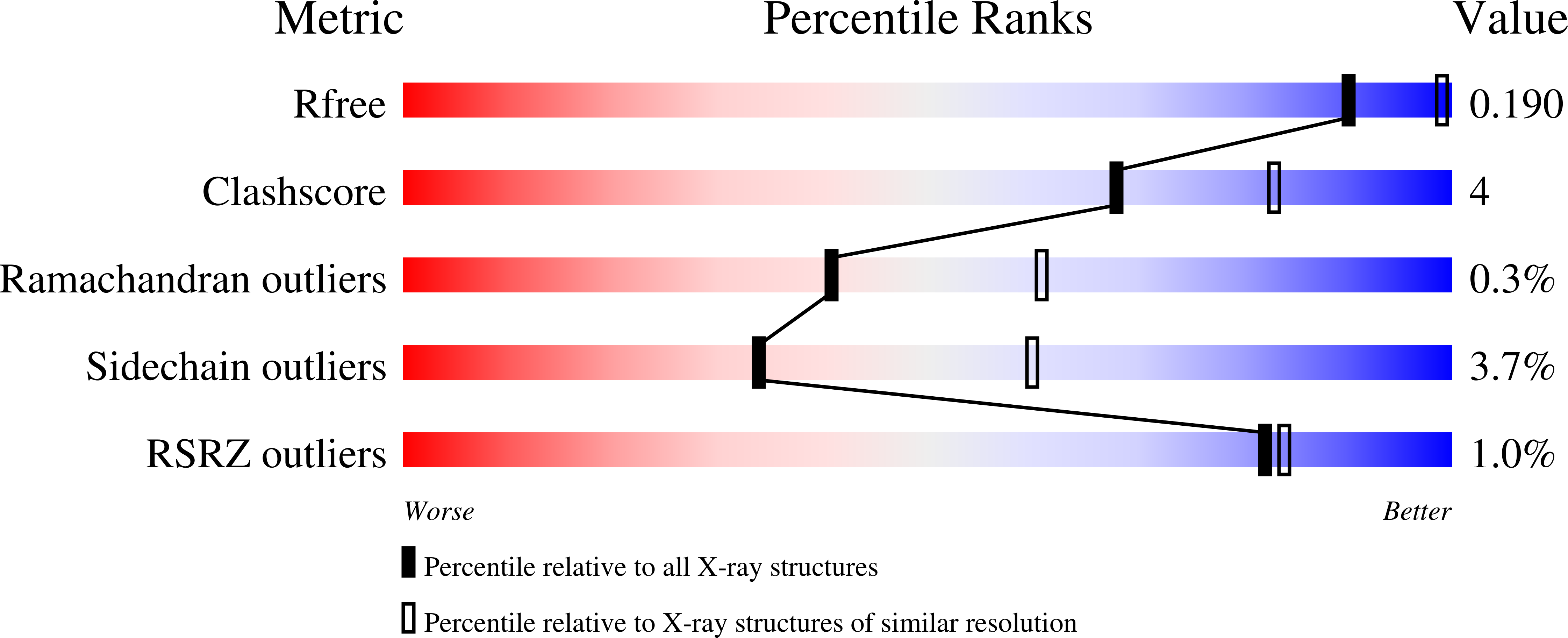
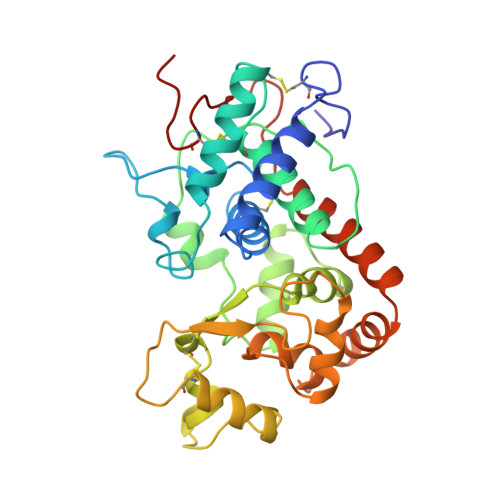
Structural analysis of the two horseradish peroxidase catalytic residue variants H42E and R38S/H42E: implications for the catalytic cycle.
Meno, K., Jennings, S., Smith, A.T., Henriksen, A., Gajhede, M.(2002) Acta Crystallogr D Biol Crystallogr 58: 1803-1812
- PubMed: 12351824
- DOI: https://doi.org/10.1107/s090744490201329x
- Primary Citation of Related Structures:
1KZM, 4ATJ - PubMed Abstract:
The crystal structures of horseradish peroxidase C (HRPC) active-site mutants H42E and R38S/H42E co-crystallized with benzhydroxamic acid (BHA) and ferulic acid (FA), respectively, have been solved. The 2.5 A crystal structure of the H42E-BHA complex reveals that the side-chain O atoms of Glu42 occupy positions that are very similar to the positions of the two side-chain N atoms of the distal histidine in the wild-type HRPC-BHA structure. The mutation disturbs the hydrogen-bonding network extending from residue 42 to the distal calcium ion and results in the absence of the water molecule that is usually ligated to this ion in plant peroxidases. Consequently, the distal calcium ion is six- rather than seven-coordinated. In the 2.0 A R38S/H42E structure the position of Glu42 is different and no FA is observed in the distal haem pocket. This is a consequence of the absence of the Arg38 side chain, which limits the flexibility of the Glu42 side chain and modulates its acidity, making it unsuitable as a general acid-base catalyst in the reaction cycle. The water ligated to the distal calcium ion is present, showing that the wild-type distal hydrogen-bonding network is preserved. These results show why a glutamic acid residue can substitute for the conserved distal histidine in HRPC and that Arg38 plays a significant role in controlling the positioning and ionization state of the residue at position 42. Furthermore, these structures indicate that changes in the distal cavity are conveyed through the distal hydrogen-bonding network to the distal calcium site.
Organizational Affiliation:
Protein Structure Group, Department of Chemistry, University of Copenhagen, Universitetsparken 5, DK-2100 Copenhagen Ø, Denmark.