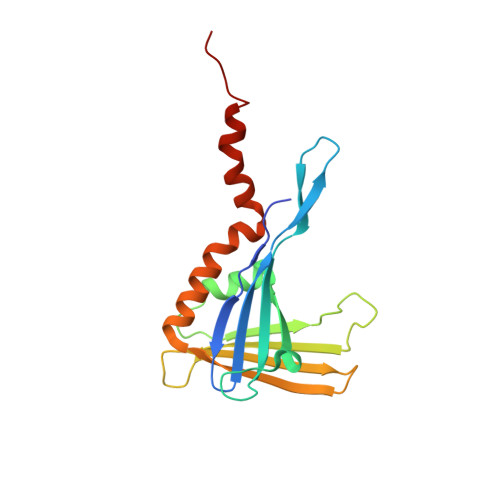
A family portrait: structural comparison of the Whirly proteins from Arabidopsis thaliana and Solanum tuberosum.
Cappadocia, L., Parent, J.S., Sygusch, J., Brisson, N.(2013) Acta Crystallogr Sect F Struct Biol Cryst Commun 69: 1207-1211
- PubMed: 24192350
- DOI: https://doi.org/10.1107/S1744309113028698
- Primary Citation of Related Structures:
4KOO, 4KOP, 4KOQ - PubMed Abstract:
DNA double-strand breaks are highly detrimental genomic lesions that routinely arise in genomes. To protect the integrity of their genetic information, all organisms have evolved specialized DNA-repair mechanisms. Whirly proteins modulate DNA repair in plant chloroplasts and mitochondria by binding single-stranded DNA in a non-sequence-specific manner. Although most of the results showing the involvement of the Whirly proteins in DNA repair have been obtained in Arabidopsis thaliana, only the crystal structures of the potato Whirly proteins WHY1 and WHY2 have been reported to date. The present report of the crystal structures of the three Whirly proteins from A. thaliana (WHY1, WHY2 and WHY3) reveals that these structurally similar proteins assemble into tetramers. Furthermore, structural alignment with a potato WHY2-DNA complex reveals that the residues in these proteins are properly oriented to bind single-stranded DNA in a non-sequence-specific manner.
Organizational Affiliation:
Department of Biochemistry, Université de Montréal, PO Box 6128, Station Centre-Ville, Montréal, Québec H3C 3J7, Canada.