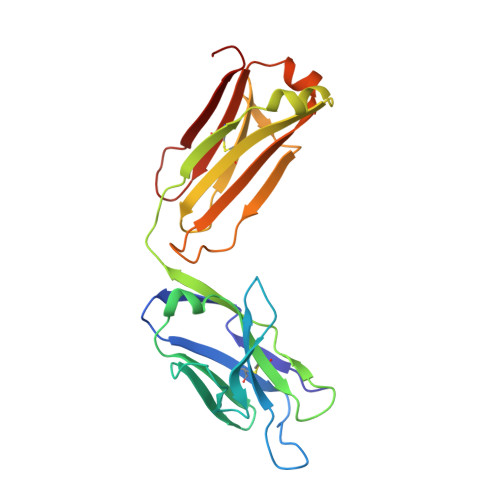
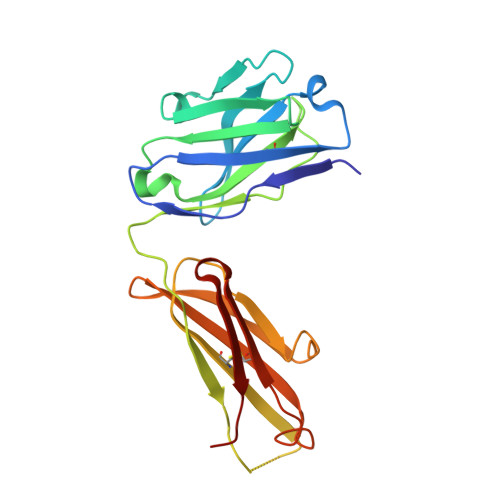
The antigen-binding site of an N-propionylated polysialic acid-specific antibody protective against group B meningococci is consistent with extended epitopes.
Johal, A.R., Jarrell, H.C., Letts, J.A., Khieu, N.H., Landry, R.C., Jachymek, W., Yang, Q., Jennings, H.J., Brisson, J.R., Evans, S.V.(2013) Glycobiology 23: 946-954
- PubMed: 23704298
- DOI: https://doi.org/10.1093/glycob/cwt031
- Primary Citation of Related Structures:
4HXA, 4HXB - PubMed Abstract:
Monoclonal antibodies 13D9 and 6B9 are both specific for N-propionylated polysialic acid (NPrPSA); however, while 13D9 is protective against meningitis caused by group B meningococci and Escherichia coli capsular type K1 infection, 6B9 is not. The crystal structures of the Fabs from the two antibodies determined at 2.06 and 2.45 Å resolutions, respectively, reveal markedly different combining sites, where only the surface of 13D9 is consistent with the recognition of extended helical epitopes known to exist in the capsular polysaccharides of etiological agents of meningitis. Interestingly, complementarity determining region H2 on 13D9 lies in a non-canonical conformation that docking studies show is a critical feature in the generation of negative free energy of binding. Finally, the model of extended NPrPSA decasaccharide bound to 13D9 derived from docking studies is consistent with saturation transfer difference nuclear magnetic resonance experiments. Together, these results provide further evidence that extended epitopes have the ability to break immune tolerance associated with the polysialic acid capsule of these pathogens.
Organizational Affiliation:
Department of Biochemistry and Microbiology, University of Victoria, Victoria, BC, Canada.