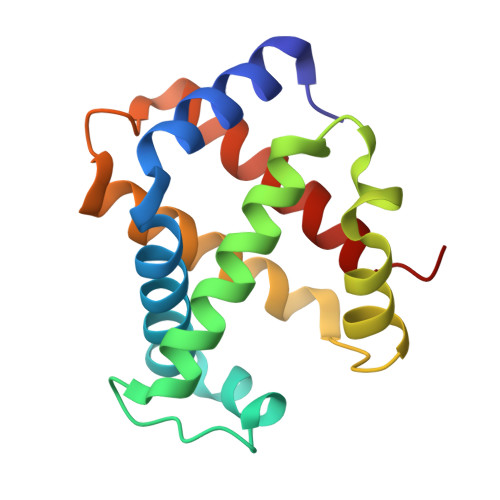
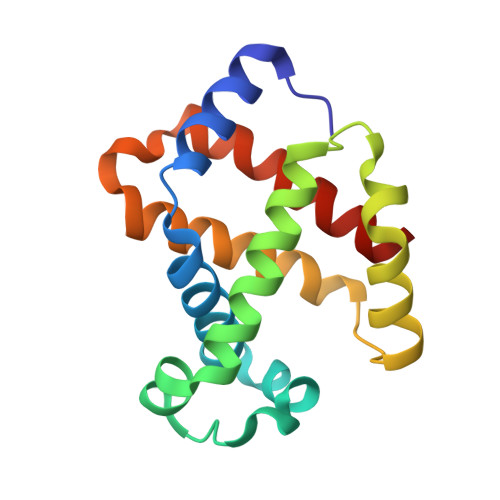
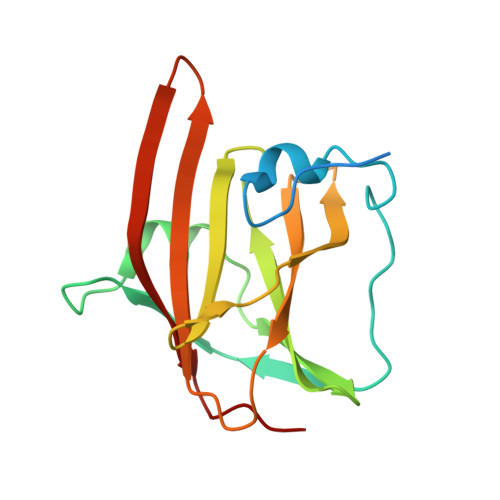
Structure of the Hemoglobin-IsdH Complex Reveals the Molecular Basis of Iron Capture by Staphylococcus aureus
Dickson, C.F., Krishna Kumar, K., Jacques, D.A., Malmirchegini, G.R., Spirig, T., Mackay, J.P., Clubb, R.T., Guss, J.M., Gell, D.A.(2014) J Biol Chem 289: 6728-6738
- PubMed: 24425866
- DOI: https://doi.org/10.1074/jbc.M113.545566
- Primary Citation of Related Structures:
4FC3, 4IJ2 - PubMed Abstract:
Staphylococcus aureus causes life-threatening disease in humans. The S. aureus surface protein iron-regulated surface determinant H (IsdH) binds to mammalian hemoglobin (Hb) and extracts heme as a source of iron, which is an essential nutrient for the bacteria. However, the process of heme transfer from Hb is poorly understood. We have determined the structure of IsdH bound to human Hb by x-ray crystallography at 4.2 Å resolution, revealing the structural basis for heme transfer. One IsdH molecule is bound to each α and β Hb subunit, suggesting that the receptor acquires iron from both chains by a similar mechanism. Remarkably, two near iron transporter (NEAT) domains in IsdH perform very different functions. An N-terminal NEAT domain binds α/β globin through a site distant from the globin heme pocket and, via an intervening structural domain, positions the C-terminal heme-binding NEAT domain perfectly for heme transfer. These data, together with a 2.3 Å resolution crystal structure of the isolated N-terminal domain bound to Hb and small-angle x-ray scattering of free IsdH, reveal how multiple domains of IsdH cooperate to strip heme from Hb. Many bacterial pathogens obtain iron from human hemoglobin using proteins that contain multiple NEAT domains and other domains whose functions are poorly understood. Our results suggest that, rather than acting as isolated units, NEAT domains may be integrated into higher order architectures that employ multiple interaction interfaces to efficiently extract heme from host proteins.
Organizational Affiliation:
Menzies Research Institute Tasmania, University of Tasmania, Hobart, Tasmania 7000, Australia.