The Natural Product Antibiotic Terreic Acid is a Mechanism-Based Inhibitor of the Bacterial Enzyme MurA in vitro but not in vivo.
Han, H., Yang, Y., Olesen, S.H., Becker, A., Betzi, S., Schonbrunn, E.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| UDP-N-acetylglucosamine 1-carboxyvinyltransferase | 419 | Escherichia coli K-12 | Mutation(s): 0 Gene Names: b3189, JW3156, murA, MurA (MurZ), murZ EC: 2.5.1.7 | 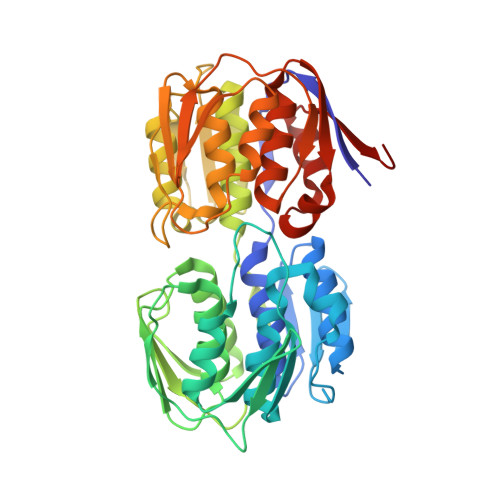 | |
UniProt | |||||
Find proteins for P0A749 (Escherichia coli (strain K12)) Explore P0A749 Go to UniProtKB: P0A749 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0A749 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| UD1 Query on UD1 | C [auth A] | URIDINE-DIPHOSPHATE-N-ACETYLGLUCOSAMINE C17 H27 N3 O17 P2 LFTYTUAZOPRMMI-CFRASDGPSA-N |  | ||
| PO4 Query on PO4 | D [auth A] | PHOSPHATE ION O4 P NBIIXXVUZAFLBC-UHFFFAOYSA-K |  | ||
| GOL Query on GOL | B [auth A] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 60.56 | α = 90 |
| b = 64.13 | β = 90 |
| c = 134.99 | γ = 90 |
| Software Name | Purpose |
|---|---|
| StructureStudio | data collection |
| CNS | refinement |
| XDS | data reduction |
| XDS | data scaling |
| CNS | phasing |