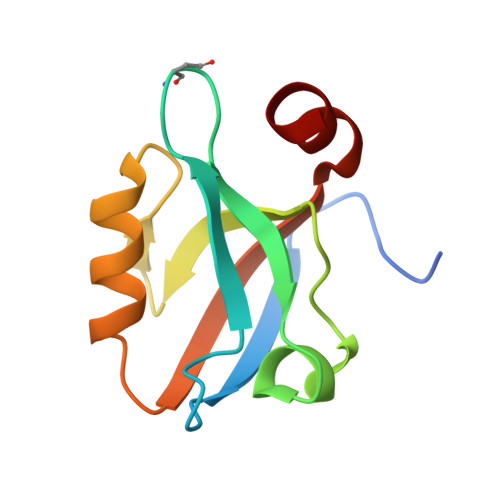
Novel conformational aspects of the third PDZ domain of the neuronal post-synaptic density-95 protein revealed from two 1.4A X-ray structures
Camara-Artigas, A., Murciano-Calles, J., Gavira, J.A., Cobos, E.S., Martinez, J.C.(2010) J Struct Biol 170: 565-569
- PubMed: 20227506
- DOI: https://doi.org/10.1016/j.jsb.2010.03.005
- Primary Citation of Related Structures:
3I4W, 3K82 - PubMed Abstract:
The crystal structure of the third PDZ domain of the neuronal post-synaptic density-95 protein (PSD95-PDZ3, residues 302-402) has been solved at 1.4 and 1.35A from two different crystal forms. These structures lack the cloning artefact present in the carboxyl terminal sequence of the former crystallographic structures and they belong to the space groups P4(3) and P1. The new PDZ structures are identical between the two crystal forms and among the four chains of the P1 crystal form. When we compare the new structures with the previous ones, some important conformational differences in the C-terminal alpha-helix and in the loop connecting beta2 and beta3 strands have been found. Additionally, the high resolution of the new structures has allowed us to indentify a succinimide residue at the position corresponding to Asp332 in the beta2-beta3 loop, which may contribute to the alternate conformation of this loop, and at the same time, to the interaction between residues from this loop and the C-terminal alpha-helix. Thus, these features would have implications in the recently proposed allosteric role of this third alpha-helix in the binding of the carboxyl terminal fragments to the PSD95-PDZ3.
Organizational Affiliation:
Department of Physical Chemistry, Biochemistry and Inorganic Chemistry, University of Almería, 04120 Almería, Spain. acamara@ual.es