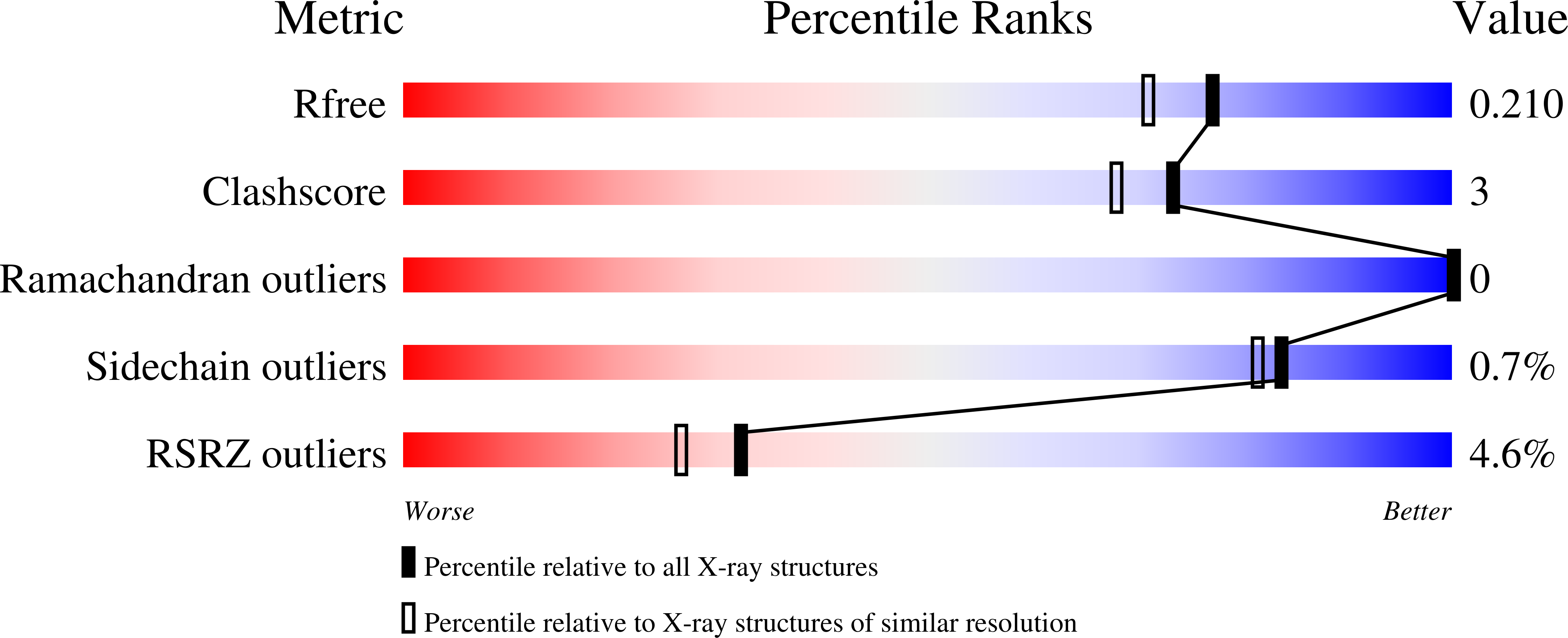
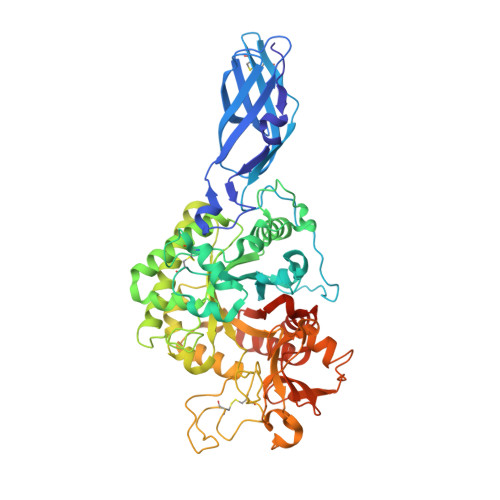
Crystal structures of Vibrio harveyi chitinase A complexed with chitooligosaccharides: implications for the catalytic mechanism
Songsiriritthigul, C., Pantoom, S., Aguda, A.H., Robinson, R.C., Suginta, W.(2008) J Struct Biol 162: 491-499
- PubMed: 18467126
- DOI: https://doi.org/10.1016/j.jsb.2008.03.008
- Primary Citation of Related Structures:
3B8S, 3B9A, 3B9D, 3B9E - PubMed Abstract:
This research describes four X-ray structures of Vibrio harveyi chitinase A and its catalytically inactive mutant (E315M) in the presence and absence of substrates. The overall structure of chitinase A is that of a typical family-18 glycosyl hydrolase comprising three distinct domains: (i) the amino-terminal chitin-binding domain; (ii) the main catalytic (alpha/beta)(8) TIM-barrel domain; and (iii) the small (alpha+beta) insertion domain. The catalytic cleft of chitinase A has a long, deep groove, which contains six chitooligosaccharide ring-binding subsites (-4)(-3)(-2)(-1)(+1)(+2). The binding cleft of the ligand-free E315M is partially blocked by the C-terminal (His)(6)-tag. Structures of E315M-chitooligosaccharide complexes display a linear conformation of pentaNAG, but a bent conformation of hexaNAG. Analysis of the final 2F(o)-F(c) omit map of E315M-NAG6 reveals the existence of the linear conformation of the hexaNAG at a lower occupancy with respect to the bent conformation. These crystallographic data provide evidence that the interacting sugars undergo conformational changes prior to hydrolysis by the wild-type enzyme.
Organizational Affiliation:
School of Biochemistry, Suranaree University of Technology, Nakhon Ratchasima 30000, Thailand.