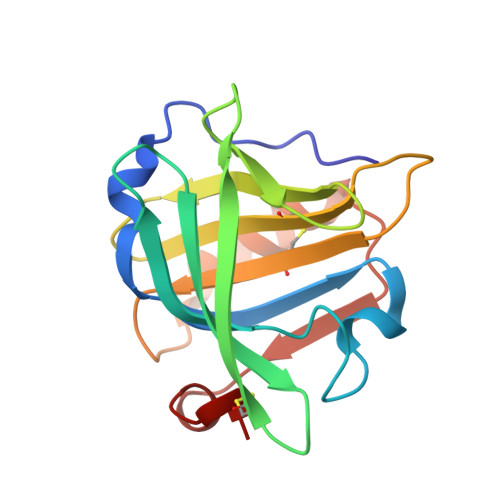
Two modes of fatty acid binding to bovine beta-lactoglobulin-crystallographic and spectroscopic studies
Loch, J.I., Polit, A., Gorecki, A., Bonarek, P., Kurpiewska, K., Dziedzicka-Wasylewska, M., Lewinski, K.(2011) J Mol Recognit 24: 341-349
- PubMed: 21360616
- DOI: https://doi.org/10.1002/jmr.1084
- Primary Citation of Related Structures:
3NPO, 3NQ3, 3NQ9 - PubMed Abstract:
Lactoglobulin is a natural protein present in bovine milk and common component of human diet, known for binding with high affinity wide range of hydrophobic compounds, among them fatty acids 12-20 carbon atoms long. Shorter fatty acids were reported as not binding to β-lactoglobulin. We used X-ray crystallography and fluorescence spectroscopy to show that lactoglobulin binds also 8- and 10-carbon caprylic and capric acids, however with lower affinity. The determined apparent association constant for lactoglobulin complex with caprylic acid is 10.8 ± 1.7 × 10(3) M(-1), while for capric acid is 6.0 ± 0.5 × 10(3) M(-1). In crystal structures determined with resolution 1.9 Å the caprylic acid is bound in upper part of central calyx near polar residues located at CD loop, while the capric acid is buried deeper in the calyx bottom and does not interact with polar residues at CD loop. In both structures, water molecule hydrogen-bonded to carboxyl group of fatty acid is observed. Different location of ligands in the binding site indicates that competition between polar and hydrophobic interactions is an important factor determining position of the ligand in β-barrel.
Organizational Affiliation:
Faculty of Chemistry, Department of Crystal Physics and Crystal Chemistry, Jagiellonian University, Ingardena 3, 30-060 Kraków, Poland.