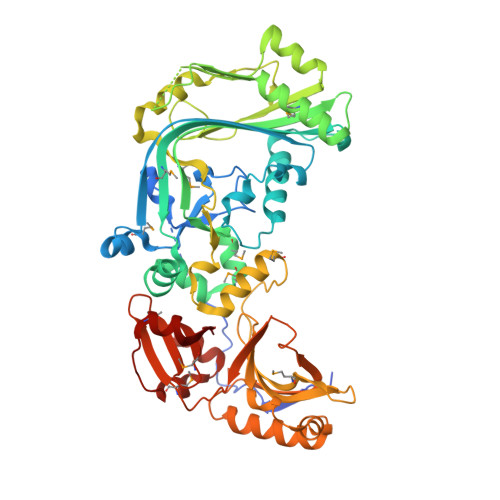
Structure of a membrane-attack complex/perforin (MACPF) family protein from the human gut symbiont Bacteroides thetaiotaomicron.
Xu, Q., Abdubek, P., Astakhova, T., Axelrod, H.L., Bakolitsa, C., Cai, X., Carlton, D., Chen, C., Chiu, H.J., Clayton, T., Das, D., Deller, M.C., Duan, L., Ellrott, K., Farr, C.L., Feuerhelm, J., Grant, J.C., Grzechnik, A., Han, G.W., Jaroszewski, L., Jin, K.K., Klock, H.E., Knuth, M.W., Kozbial, P., Krishna, S.S., Kumar, A., Lam, W.W., Marciano, D., Miller, M.D., Morse, A.T., Nigoghossian, E., Nopakun, A., Okach, L., Puckett, C., Reyes, R., Tien, H.J., Trame, C.B., van den Bedem, H., Weekes, D., Wooten, T., Yeh, A., Zhou, J., Hodgson, K.O., Wooley, J., Elsliger, M.A., Deacon, A.M., Godzik, A., Lesley, S.A., Wilson, I.A.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 1297-1305
- PubMed: 20944225
- DOI: https://doi.org/10.1107/S1744309110023055
- Primary Citation of Related Structures:
3KK7 - PubMed Abstract:
Membrane-attack complex/perforin (MACPF) proteins are transmembrane pore-forming proteins that are important in both human immunity and the virulence of pathogens. Bacterial MACPFs are found in diverse bacterial species, including most human gut-associated Bacteroides species. The crystal structure of a bacterial MACPF-domain-containing protein BT_3439 (Bth-MACPF) from B. thetaiotaomicron, a predominant member of the mammalian intestinal microbiota, has been determined. Bth-MACPF contains a membrane-attack complex/perforin (MACPF) domain and two novel C-terminal domains that resemble ribonuclease H and interleukin 8, respectively. The entire protein adopts a flat crescent shape, characteristic of other MACPF proteins, that may be important for oligomerization. This Bth-MACPF structure provides new features and insights not observed in two previous MACPF structures. Genomic context analysis infers that Bth-MACPF may be involved in a novel protein-transport or nutrient-uptake system, suggesting an important role for these MACPF proteins, which were likely to have been inherited from eukaryotes via horizontal gene transfer, in the adaptation of commensal bacteria to the host environment.
Organizational Affiliation:
Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, Menlo Park, CA, USA.