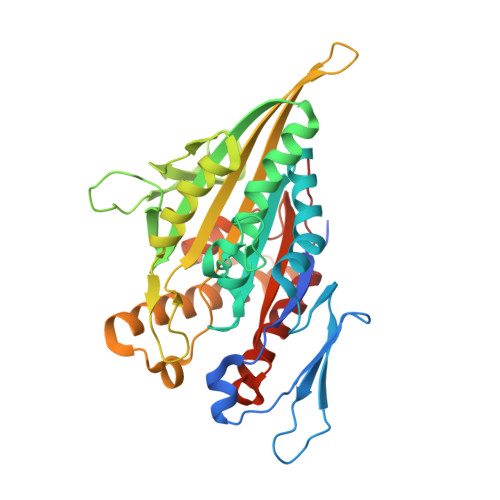
ATP Hydrolysis in Eg5 Kinesin Involves a Catalytic Two-water Mechanism.
Parke, C.L., Wojcik, E.J., Kim, S., Worthylake, D.K.(2010) J Biol Chem 285: 5859-5867
- PubMed: 20018897
- DOI: https://doi.org/10.1074/jbc.M109.071233
- Primary Citation of Related Structures:
3HQD - PubMed Abstract:
Motor proteins couple steps in ATP binding and hydrolysis to conformational switching both in and remote from the active site. In our kinesin.AMPPPNP crystal structure, closure of the active site results in structural transformations appropriate for microtubule binding and organizes an orthosteric two-water cluster. We conclude that a proton is shared between the lytic water, positioned for gamma-phosphate attack, and a second water that serves as a general base. To our knowledge, this is the first experimental detection of the catalytic base for any ATPase. Deprotonation of the second water by switch residues likely triggers subsequent large scale structural rearrangements. Therefore, the catalytic base is responsible for initiating nucleophilic attack of ATP and for relaying the positive charge over long distances to initiate mechanotransduction. Coordination of switch movements via sequential proton transfer along paired water clusters may be universal for nucleotide triphosphatases with conserved active sites, such as myosins and G-proteins.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, Louisiana State University Health Sciences Center, New Orleans, Louisiana 70112, USA.