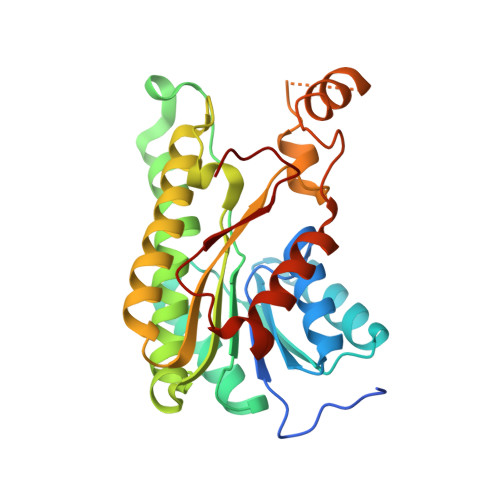
Inhibition kinetics and emodin cocrystal structure of a type II polyketide ketoreductase
Korman, T.P., Tan, Y.H., Wong, J., Luo, R., Tsai, S.-C.(2008) Biochemistry 47: 1837-1847
- PubMed: 18205400
- DOI: https://doi.org/10.1021/bi7016427
- Primary Citation of Related Structures:
2RH4, 2RHC, 2RHR - PubMed Abstract:
Type II polyketides are a class of natural products that include pharmaceutically important aromatic compounds such as the antibiotic tetracycline and antitumor compound doxorubicin. The type II polyketide synthase (PKS) is a complex consisting of 5-10 standalone domains homologous to fatty acid synthase (FAS). Polyketide ketoreductase (KR) provides regio- and stereochemical diversity during the reduction. How the type II polyketide KR specifically reduces only the C9 carbonyl group is not well understood. The cocrystal structures of actinorhodin polyketide ketoreductase (actKR) bound with NADPH or NADP+ and the inhibitor emodin were solved with the wild type and P94L mutant of actKR, revealing the first observation of a bent p-quinone in an enzyme active site. Molecular dynamics simulation help explain the origin of the bent geometry. Extensive screening for in vitro substrates shows that unlike FAS KR, the actKR prefers bicyclic substrates. Inhibition kinetics indicate that actKR follows an ordered Bi Bi mechanism. Together with docking simulations that identified a potential phosphopantetheine binding groove, the structural and functional studies reveal that the C9 specificity is a result of active site geometry and substrate ring constraints. The results lay the foundation for the design of novel aromatic polyketide natural products with different reduction patterns.
Organizational Affiliation:
Department of Molecular Biology and Biochemistry, University of California, Irvine, California 92697, USA.