4-Aminophenylalanine and 4-aminocyclohexylalanine derivatives as potent, selective, and orally bioavailable inhibitors of dipeptidyl peptidase IV.
Duffy, J.L., Kirk, B.A., Wang, L., Eiermann, G.J., He, H., Leiting, B., Lyons, K.A., Patel, R.A., Patel, S.B., Petrov, A., Scapin, G., Wu, J.K., Thornberry, N.A., Weber, A.E.(2007) Bioorg Med Chem Lett 17: 2879-2885
- PubMed: 17350841
- DOI: https://doi.org/10.1016/j.bmcl.2007.02.066
- Primary Citation of Related Structures:
2OPH - PubMed Abstract:
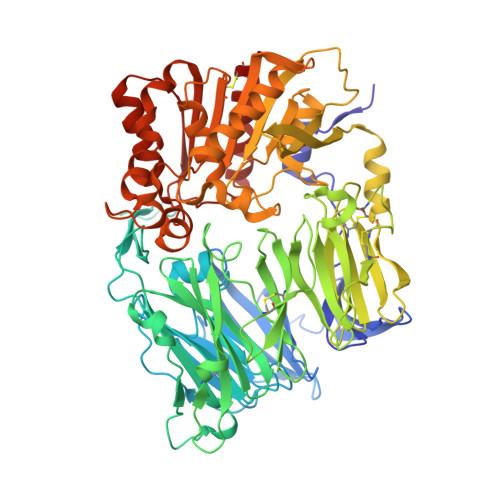
A novel series of 4-aminophenylalanine and 4-aminocyclohexylalanine derivatives were designed and evaluated as inhibitors of dipeptidyl peptidase IV (DPP-4). The phenylalanine series afforded compounds such as 10 that were potent and selective (DPP-4, IC(50)=28nM), but exhibited limited oral bioavailability. The corresponding cyclohexylalanine derivatives such as 25 afforded improved PK exposure and efficacy in a murine OGTT experiment. The X-ray crystal structure of 25 bound to the DPP-4 active site is presented.
Organizational Affiliation:
Department of Medicinal Chemistry, Merck Research Laboratories, Rahway, NJ 07065, USA. joseph_duffy@merck.com