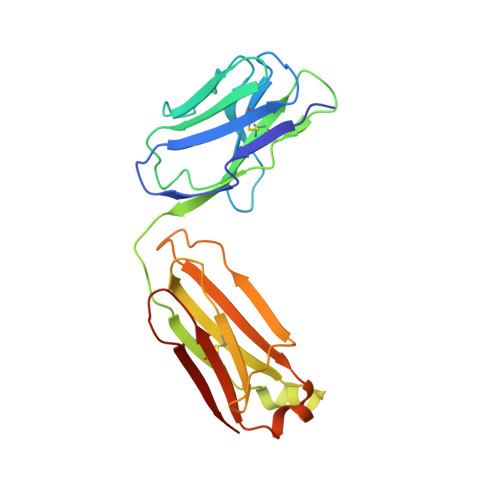
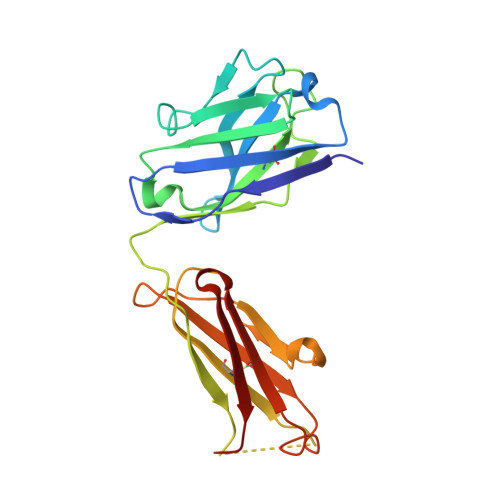
Interfacial metal and antibody recognition.
Zhou, T., Hamer, D.H., Hendrickson, W.A., Sattentau, Q.J., Kwong, P.D.(2005) Proc Natl Acad Sci U S A 102: 14575-14580
- PubMed: 16195378
- DOI: https://doi.org/10.1073/pnas.0507267102
- Primary Citation of Related Structures:
2ADG, 2ADI, 2ADJ - PubMed Abstract:
The unique ligation properties of metal ions are widely exploited by proteins, with approximately one-third of all proteins estimated to be metalloproteins. Although antibodies use various mechanisms for recognition, to our knowledge, none has ever been characterized that uses an interfacial metal. We previously described a family of CD4-reactive antibodies, the archetype being Q425. CD4:Q425 engagement does not interfere with CD4:HIV-1 gp120 envelope glycoprotein binding, but it blocks subsequent steps required for viral entry. Here, we use surface-plasmon resonance to show that Q425 requires calcium for recognition of CD4. Specifically, Q425 binding of calcium resulted in a 55,000-fold enhancement in affinity for CD4. X-ray crystallographic analyses of Q425 in the presence of Ca(2+), Ba(2+), or EDTA revealed an exposed metal-binding site, partially coordinated by five atoms contributed from four antibody complementarity-determining regions. The results suggest that Q425 recognition of CD4 involves direct ligation of antigen by the Q425-held calcium, with calcium binding each ligating atom of CD4 with approximately 1.5 kcal/mol of binding energy. This energetic contribution, which is greater than that from a typical protein atom, demonstrates how interfacial metal ligation can play a unique role in antigen recognition.
Organizational Affiliation:
Vaccine Research Center, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD 20892, USA.