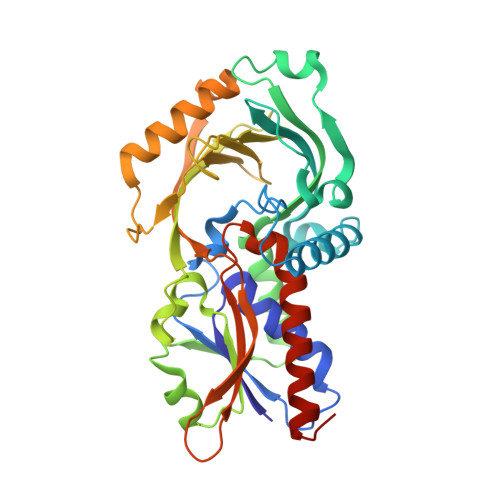
Crystal structure of human D-amino acid oxidase: Context-dependent variability of the backbone conformation of the VAAGL hydrophobic stretch located at the si-face of the flavin ring
Kawazoe, T., Tsuge, H., Pilone, M.S., Fukui, K.(2006) Protein Sci 15: 2708-2717
- PubMed: 17088322
- DOI: https://doi.org/10.1110/ps.062421606
- Primary Citation of Related Structures:
2DU8 - PubMed Abstract:
In the brain, the extensively studied FAD-dependent enzyme D-amino acid oxidase (DAO) degrades the gliotransmitter D-serine, a potent activator of N-methyl-D-aspartate type glutamate receptors, and evidence suggests that DAO, together with its activator G72 protein, may play a key role in the pathophysiology of schizophrenia. Indeed, its potential clinical importance highlights the need for structural and functional analyses of human DAO. We recently succeeded in purifying human DAO, and found that it weakly binds FAD and shows a significant slower rate of flavin reduction compared with porcine DAO. However, the molecular basis for the different kinetic features remains unclear because the active site of human DAO was considered to be virtually identical to that of porcine DAO, as would be expected from the 85% sequence identity. To address this issue, we determined the crystal structure of human DAO in complex with a competitive inhibitor benzoate, at a resolution of 2.5 Angstrom. The overall dimeric structure of human DAO is similar to porcine DAO, and the catalytic residues are fully conserved at the re-face of the flavin ring. However, at the si-face of the flavin ring, despite the strict sequence identity, a hydrophobic stretch (residues 47-51, VAAGL) exists in a significantly different conformation compared with both of the independently determined porcine DAO-benzoate structures. This suggests that a context-dependent conformational variability of the hydrophobic stretch accounts for the low affinity for FAD as well as the slower rate of flavin reduction, thus highlighting the unique features of the human enzyme.
Organizational Affiliation:
The Institute for Enzyme Research, The University of Tokushima, 3-18-15 Kuramoto, Tokushima 770-8503, Japan.