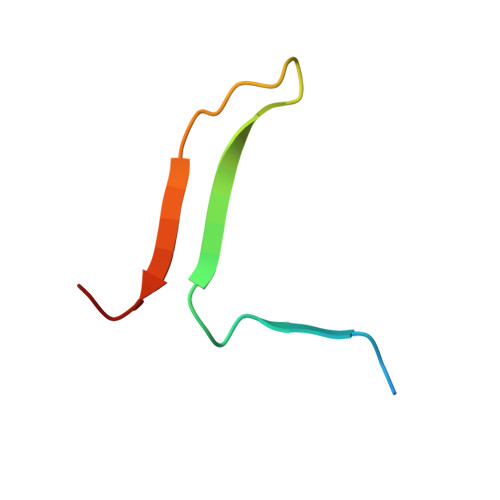
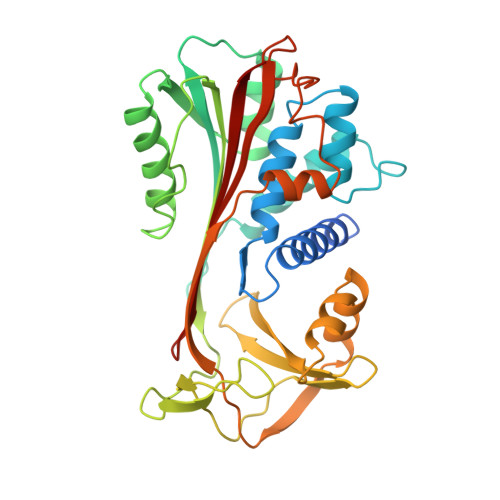
Crystal structure of cleaved human alpha 1-antichymotrypsin at 2.7 A resolution and its comparison with other serpins.
Baumann, U., Huber, R., Bode, W., Grosse, D., Lesjak, M., Laurell, C.B.(1991) J Mol Biol 218: 595-606
- PubMed: 2016749
- DOI: https://doi.org/10.1016/0022-2836(91)90704-a
- Primary Citation of Related Structures:
2ACH - PubMed Abstract:
The crystal structure of proteolytically modified human alpha 1-antichymotrypsin (ACT), a member of the serpin superfamily, has been solved by Paterson search techniques and refined to an R-factor of 18.0% at 2.7 A resolution with mean deviations from standard bond lengths and angles of 0.013 A and 3.1 degrees, respectively. The final model consists of 374 amino acid residues, 126 solvent molecules and five sugar residues. Asn70 could be identified unambiguously as a glycosylation site and Asn104 is probably also glycosylated. The structure of cleaved ACT is compared with cleaved alpha 1-antitrypsin (alpha 1 PI) and with plakalbumin, which are prototypical models for cleaved and intact serpins, respectively. Cleaved ACT is very similar to cleaved alpha 1 PI; in particular, it has strand s4A, which is liberated by proteolysis, inserted as the middle strand in beta-sheet A. ACT and alpha 1 PI differ locally only at sites of insertions, except at the segment s3C-turn-s4C, which is displaced by several angström units. This region of ACT is involved in DNA binding.
Organizational Affiliation:
Max-Planck-Institut für Biochemie, Martinsried, Germany.