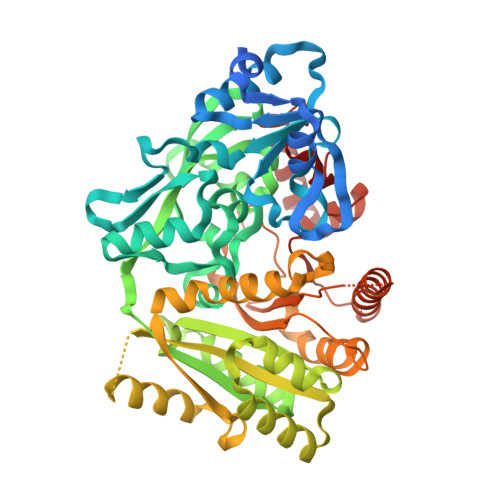
Structures of Michaelis and Product Complexes of Plant Cytokinin Dehydrogenase: Implications for Flavoenzyme Catalysis
Malito, E., Coda, A., Bilyeu, K., Fraaije, M.W., Mattevi, A.(2004) J Mol Biol 341: 1237
- PubMed: 15321719
- DOI: https://doi.org/10.1016/j.jmb.2004.06.083
- Primary Citation of Related Structures:
1W1O, 1W1Q, 1W1R, 1W1S - PubMed Abstract:
Cytokinins form a diverse class of compounds that are essential for plant growth. Cytokinin dehydrogenase has a major role in the control of the levels of these plant hormones by catalysing their irreversible oxidation. The crystal structure of Zea mays cytokinin dehydrogenase displays the same two-domain topology of the flavoenzymes of the vanillyl-alcohol oxidase family but its active site cannot be related to that of any other family member. The X-ray analysis reveals a bipartite architecture of the catalytic centre, which consists of a funnel-shaped region on the protein surface and an internal cavity lined by the flavin ring. A pore with diameter of about 4A connects the two active-site regions. Snapshots of two critical steps along the reaction cycle were obtained through the structural analysis of the complexes with a slowly reacting substrate and the reaction product, which correspond to the states immediately before (Michaelis complex) and after (product complex) oxidation has taken place. The substrate displays a "plug-into-socket" binding mode that seals the catalytic site and precisely positions the carbon atom undergoing oxidation in close contact with the reactive locus of the flavin. A polarising H-bond between the substrate amine group and an Asp-Glu pair may facilitate oxidation. Substrate to product conversion results in small atomic movements, which lead to a planar conformation of the reaction product allowing double-bond conjugation. These features in the mechanism of amine recognition and oxidation differ from those observed in other flavin-dependent amine oxidases.
Organizational Affiliation:
Department of Genetics and Microbiology, University of Pavia, Via Ferrata 1, 27100 Pavia, Italy.