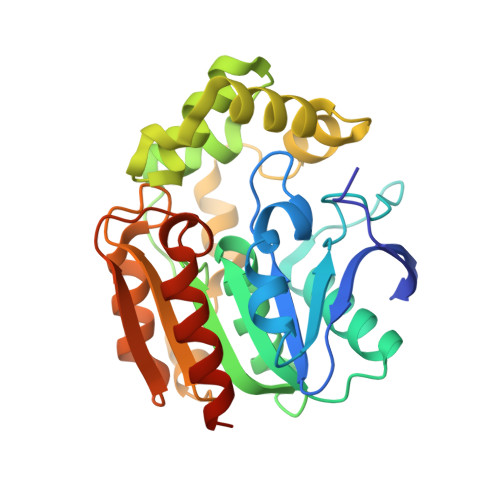
A Series of Crystal Structures of a meta-Cleavage Product Hydrolase from Pseudomonas fluorescens IP01 (CumD) Complexed with Various Cleavage Products
Fushinobu, S., Jun, S.-Y., Hidaka, M., Nojiri, H., Yamane, H., Shoun, H., Omori, T., Wakagi, T.(2005) Biosci Biotechnol Biochem 69: 491-498
- PubMed: 15784976
- DOI: https://doi.org/10.1271/bbb.69.491
- Primary Citation of Related Structures:
1UK6, 1UK7, 1UK8, 1UK9, 1UKA, 1UKB - PubMed Abstract:
Meta-cleavage product hydrolase (MCP-hydrolase) is one of the key enzymes in the microbial degradation of aromatic compounds. MCP-hydrolase produces 2-hydroxypenta-2,4-dienoate and various organic acids, according to the C6 substituent of the substrate. Comprehensive analysis of the substrate specificity of the MCP-hydrolase from Pseudomonas fluorescens IP01 (CumD) was carried out by determining the kinetic parameters for nine substrates and crystal structures complexed with eight cleavage products. CumD preferred substrates with long non-branched C6 substituents, but did not effectively hydrolyze a substrate with a phenyl group. Superimposition of the complex structures indicated that benzoate was bound in a significantly different direction than other aliphatic cleavage products. The directions of the bound organic acids appeared to be related with the k(cat) values of the corresponding substrates. The Ile139 and Trp143 residues on helix alpha4 appeared to cause steric hindrance with the aromatic ring of the substrate, which hampers base-catalyzed attack by water.
Organizational Affiliation:
Department of Biotechnology, The University of Tokyo, Japan. asfushi@mail.ecc.u-tokyo.ac.jp