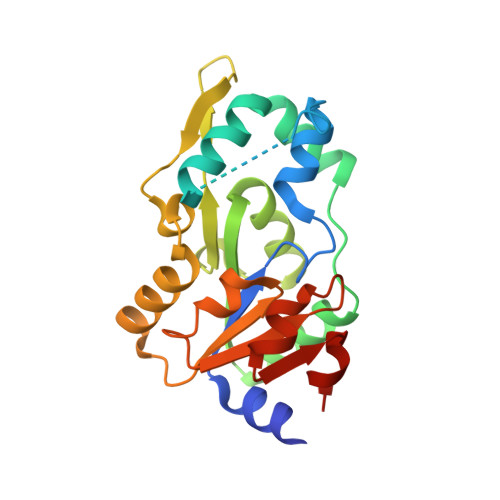
High-resolution structure of human phosphoserine phosphatase in open conformation.
Peeraer, Y., Rabijns, A., Verboven, C., Collet, J.F., Van Schaftingen, E., De Ranter, C.(2003) Acta Crystallogr D Biol Crystallogr 59: 971-977
- PubMed: 12777757
- DOI: https://doi.org/10.1107/s0907444903005407
- Primary Citation of Related Structures:
1NNL - PubMed Abstract:
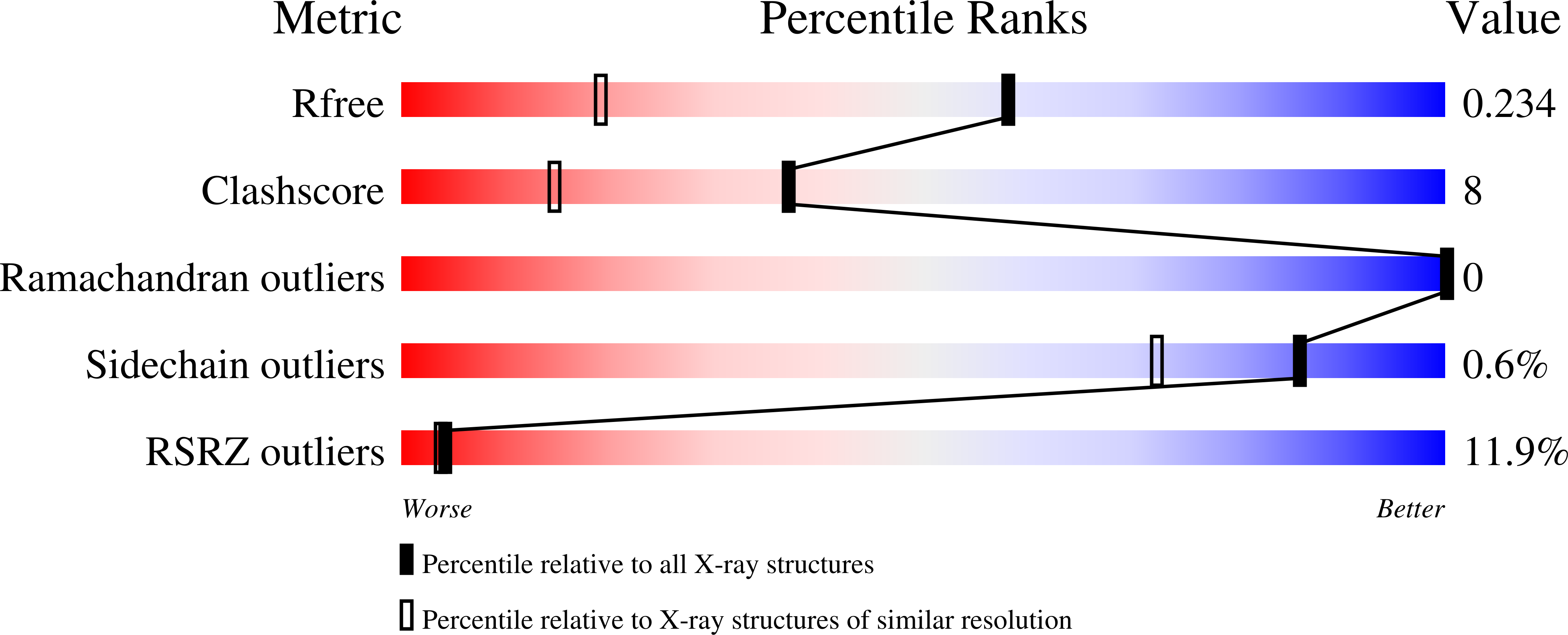
The crystal structure of human phosphoserine phosphatase (HPSP) in the open conformation has been determined at a resolution of 1.53 A. The crystals are orthorhombic, belonging to space group C222(1), with unit-cell parameters a = 49.03, b = 130.25, c = 157.29 A. The asymmetric unit contains two molecules. Phase information was derived from a multiwavelength anomalous dispersion (MAD) experiment conducted at three wavelengths using a selenomethionine-derivative crystal of HPSP. The structure was refined using CNS to a final crystallographic R value of 21.6% (R(free) = 23.4%). HPSP is a dimeric enzyme responsible for the third and final step of the l-serine biosynthesis pathway. It catalyses the Mg2+-dependent hydrolysis of l-phosphoserine. Recently, the structure of HPSP in complex with an inhibitor bound to the active site has been reported to be the open conformation of the enzyme. Here, the structure of HPSP is reported in the absence of substrate in the active site. Evidence is presented that HPSP in an uncomplexed form is in an even more open conformation than in the inhibitor complex. In this state, the enzyme is partially unfolded to allow the substrate to enter the active site. Binding of the substrate causes HPSP to shift to the closed conformation by stabilizing the partially unfolded region. In the present structure a Ca2+ ion is bound to the active site and an explanation is given why HPSP is not active when in the active site Mg2+ is replaced by a Ca2+ ion.
Organizational Affiliation:
Laboratory for Analytical Chemistry and Medicinal Physicochemistry, Faculty of Pharmaceutical Sciences, K.U. Leuven, E. Van Evenstraat 4, B-3000 Leuven, Belgium.