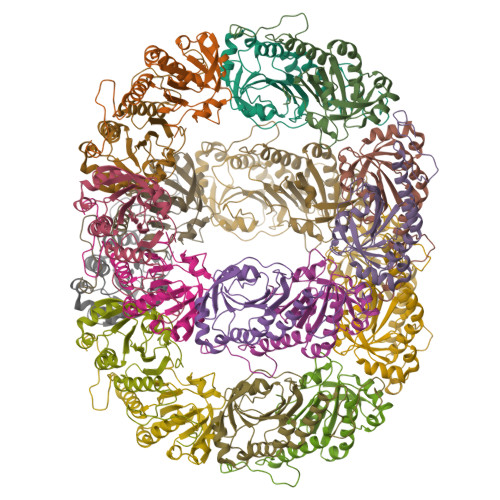
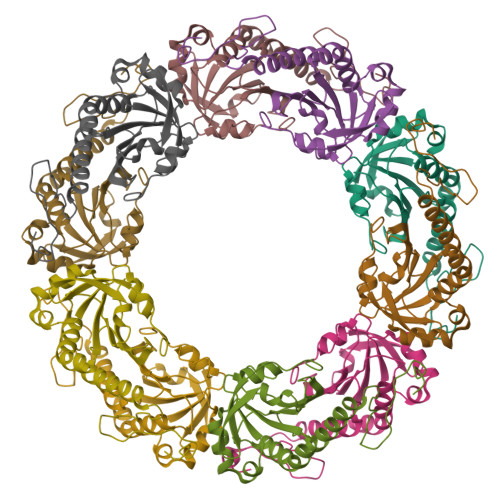
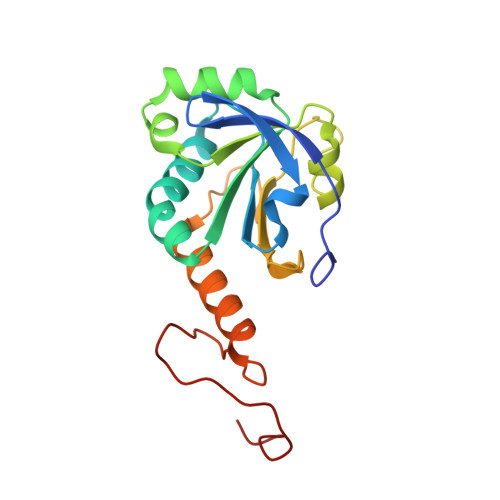
Peroxiredoxin Evolution and the Regulation of Hydrogen Peroxide Signaling
Wood, Z.A., Poole, L.B., Karplus, P.A.(2003) Science 300: 650-653
- PubMed: 12714747
- DOI: https://doi.org/10.1126/science.1080405
- Primary Citation of Related Structures:
1N8J - PubMed Abstract:
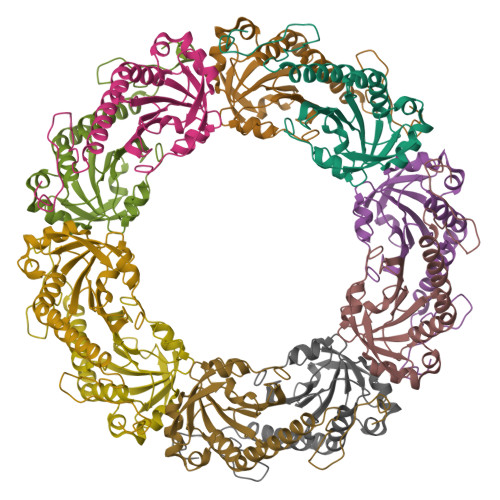
Eukaryotic 2-Cys peroxiredoxins (2-Cys Prxs) not only act as antioxidants, but also appear to regulate hydrogen peroxide-mediated signal transduction. We show that bacterial 2-Cys Prxs are much less sensitive to oxidative inactivation than are eukaryotic 2-Cys Prxs. By identifying two sequence motifs unique to the sensitive 2-Cys Prxs and comparing the crystal structure of a bacterial 2-Cys Prx at 2.2 angstrom resolution with other Prx structures, we define the structural origins of sensitivity. We suggest this adaptation allows 2-Cys Prxs to act as floodgates, keeping resting levels of hydrogen peroxide low, while permitting higher levels during signal transduction.
Organizational Affiliation:
Department of Biochemistry and Biophysics, Oregon State University, Corvallis, OR 97333, USA.