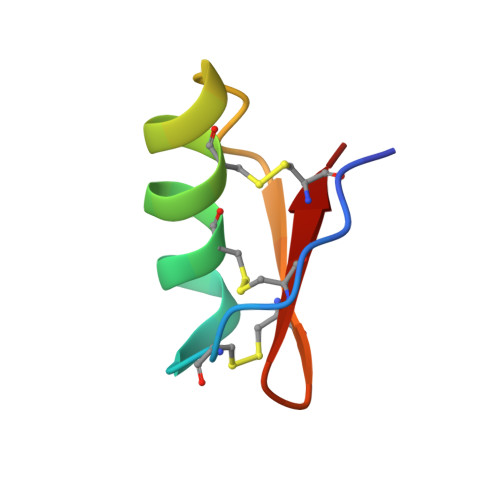
The solution structure of BmTx3B, a member of the scorpion toxin subfamily alpha-KTx 16
Wang, Y., Chen, X., Zhang, N., Wu, G., Wu, H.(2005) Proteins 58: 489-497
- PubMed: 15558557
- DOI: https://doi.org/10.1002/prot.20322
- Primary Citation of Related Structures:
1M2S - PubMed Abstract:
This article reports the solution structure of BmTx3B (alpha-KTx16.2), a potassium channel blocker belonging to the subfamily alpha-KTx16, purified from the venom of the Chinese scorpion Buthus martensi Karsch. In solution, BmTx3B assumes a typical CSalphabeta motif, with an alpha-helix connected to a triple-stranded beta-sheet by 3 disulfide bridges, which belongs to the first structural group of short-chain scorpion toxins. On the other hand, BmTx3B is quite different from other toxins (such as ChTx and AgTx2) of this group in terms of the electrostatic and hydrophobic surface distribution. The functional surface (beta-face) of the molecule is characterized by less basic residues (only 2: Lys28 and Arg35) and extra aromatic residues (Phe1, Phe9, Trp15, and Tyr37). The peptide shows a great preference for the Kca1.1 channel over the Kv channel (about a 10(3)-fold difference). The model of BmTx3B/Kca1.1 channel complex generated by docking and dynamic simulation reveals that the stable binding between the BmTx3B and Kca1.1 channel is favored by a number of aromatic pi-pi stacking interactions. The influences of these structural features on the kinetic behavior of the toxin binding to Kca1.1 channel are also discussed.
Organizational Affiliation:
State Key Laboratory of Bio-Organic and Natural Products Chemistry, Shanghai Institute of Organic Chemistry, Chinese Academy of Sciences, Shanghai, PR China.