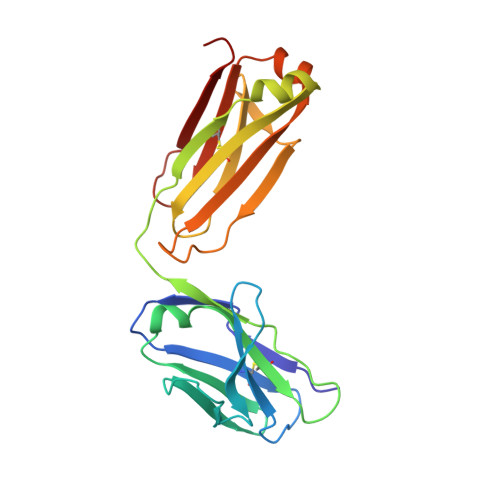
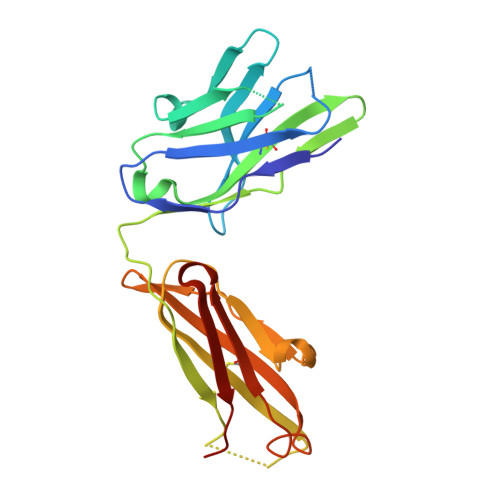
Crystal structure of a recombinant anti-estradiol Fab fragment in complex with 17beta -estradiol.
Lamminmaki, U., Kankare, J.A.(2001) J Biol Chem 276: 36687-36694
- PubMed: 11451948
- DOI: https://doi.org/10.1074/jbc.M102367200
- Primary Citation of Related Structures:
1JGL, 1JHK - PubMed Abstract:
The crystal structure of a Fab fragment of an anti-17beta-estradiol antibody 57-2 was determined in the absence and presence of the steroid ligand, 17beta-estradiol (E2), at 2.5 and 2.15-A resolutions, respectively. The antibody binds the steroid in a deep hydrophobic pocket formed at the interface between the variable domains. No major structural rearrangements take place upon ligand binding; however, a large part of the heavy chain variable domain near the binding pocket is unusually flexible and is partly stabilized when the steroid is bound. The nonpolar steroid skeleton of E2 is recognized by a number of hydrophobic interactions, whereas the two hydroxyl groups of E2 are hydrogen-bonded to the protein. Especially, the 17-hydroxyl group of E2 is recognized by an intricate hydrogen bonding network in which the 17-hydroxyl itself forms a rare four-center hydrogen bond with three polar amino acids; this hydrogen bonding arrangement accounts for the low cross-reactivity of the antibody with other estrogens such as estrone. The CDRH3 loop plays a prominent role in ligand binding. All the complementarity-determining regions of the light chain make direct contacts with the steroid, even CDRL2, which is rarely directly involved in the binding of haptens.
Organizational Affiliation:
Department of Biotechnology, University of Turku, Tykistökatu 6, 6th floor, 20520 Turku, Finland. urpo.lamminmaki@utu.fi