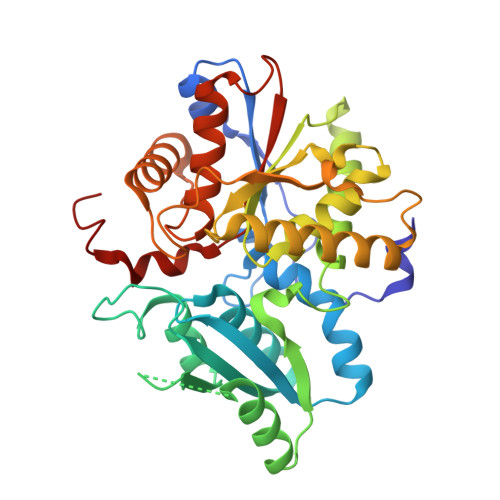
Structural Analysis of Pseudomonas 1-Aminocyclopropane-1-carboxylate Deaminase Complexes: Insight into the Mechanism of a Unique Pyridoxal-5'-phosphate Dependent Cyclopropane Ring-Opening Reaction
Karthikeyan, S., Zhou, Q., Zhao, Z., Kao, C.L., Tao, Z., Robinson, H., Liu, H.W., Zhang, H.(2004) Biochemistry 43: 13328-13339
- PubMed: 15491139
- DOI: https://doi.org/10.1021/bi048878g
- Primary Citation of Related Structures:
1TYZ, 1TZ2, 1TZJ, 1TZK, 1TZM - PubMed Abstract:
1-Aminocyclopropane-1-carboxylate (ACC) deaminase is a pyridoxal 5'-phosphate (PLP) dependent enzyme catalyzing the opening of the cyclopropane ring of ACC to give alpha-ketobutyric acid and ammonia as the products. This ring cleavage reaction is unusual because the substrate, ACC, contains no abstractable alpha-proton and the carboxyl group is retained in the product. How the reaction is initiated to generate an alpha-carbanionic intermediate, which is the common entry for most PLP-dependent reactions, is not obvious. To gain insight into this unusual ring-opening reaction, we have solved the crystal structures of ACC deaminase from Pseudomonas sp. ACP in complex with substrate ACC, an inhibitor, 1-aminocyclopropane-1-phosphonate (ACP), the product alpha-ketobutyrate, and two d-amino acids. Several notable observations of these structural studies include the following: (1) a typically elusive gem-diamine intermediate is trapped in the enzyme complex with ACC or ACP; (2) Tyr294 is in close proximity (3.0 A) to the pro-S methylene carbon of ACC in the gem-diamine complexes, implicating a direct role of this residue in the ring-opening reaction; (3) Tyr294 may also be responsible for the abstraction of the alpha-proton from d-amino acids, a prelude to the subsequent deamination reaction; (4) the steric hindrance precludes accessibility of active site functional groups to the l-amino acid substrates and may account for the stereospecificity of this enzyme toward d-amino acids. These structural data provide evidence favoring a mechanism in which the ring cleavage is induced by a nucleophilic attack at the pro-S beta-methylene carbon of ACC, with Tyr294 as the nucleophile. However, these observations are also consistent with an alternative mechanistic possibility in which the ring opening is acid-catalyzed and may be facilitated by charge relay through PLP, where Tyr294 functions as a general acid. The results of mutagenesis studies corroborated the assigned critical role for Tyr294 in the catalysis.
Organizational Affiliation:
Department of Biochemistry, University of Texas Southwestern Medical Center, Dallas, Texas 75390-9038, USA.