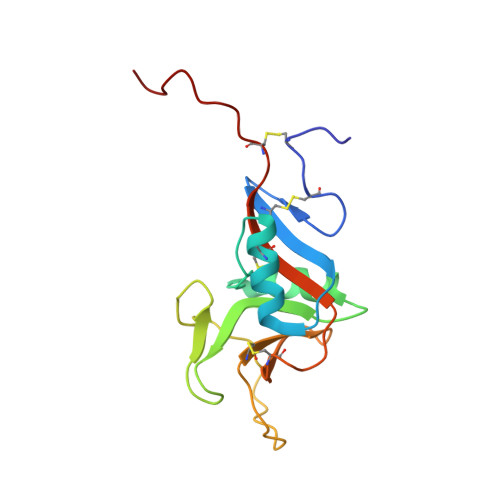
The structure of human CD23 and its interactions with IgE and CD21
Hibbert, R.G., Teriete, P., Grundy, G.J., Beavil, R.L., Reljic, R., Holers, V.M., Hannan, J.P., Sutton, B.J., Gould, H.J., McDonnell, J.M.(2005) J Exp Med 202: 751-760
- PubMed: 16172256
- DOI: https://doi.org/10.1084/jem.20050811
- Primary Citation of Related Structures:
1T8C, 1T8D - PubMed Abstract:
The low-affinity immunoglobulin E (IgE) receptor, CD23 (FcepsilonRII), binds both IgE and CD21 and, through these interactions, regulates the synthesis of IgE, the antibody isotype that mediates the allergic response. We have determined the three-dimensional structure of the C-type lectin domain of CD23 in solution by nuclear magnetic resonance spectroscopy. An analysis of concentration-dependent chemical shift perturbations have allowed us to identify the residues engaged in self-association to the trimeric state, whereas ligand-induced changes have defined the binding sites for IgE and CD21. The results further reveal that CD23 can bind both ligands simultaneously. Despite the C-type lectin domain structure, none of the interactions require calcium. We also find that IgE and CD23 can interact to form high molecular mass multimeric complexes. The interactions that we have described provide a solution to the paradox that CD23 is involved in both up- and down-regulation of IgE and provide a structural basis for the development of inhibitors of allergic disease.
Organizational Affiliation:
Laboratory of Molecular Biophysics, Department of Biochemistry, University of Oxford, Oxford OX1 3QU, England, UK.