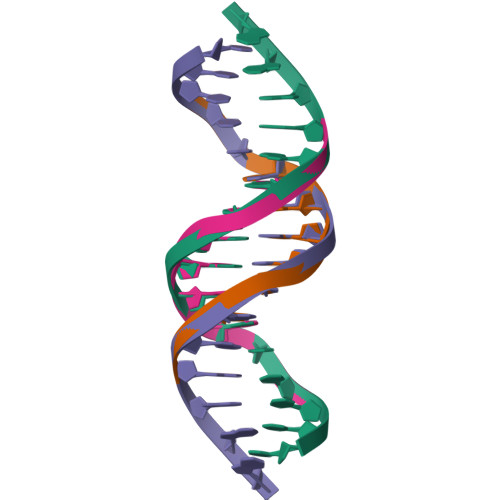
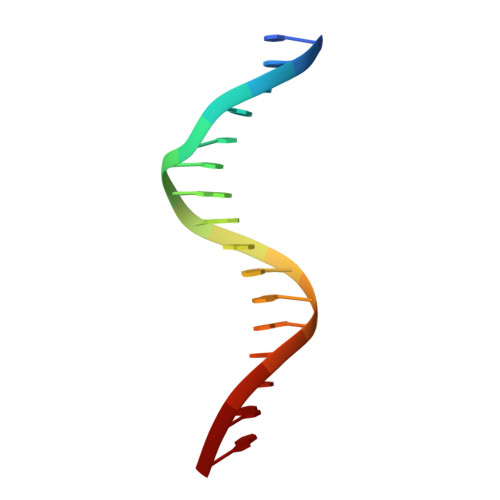
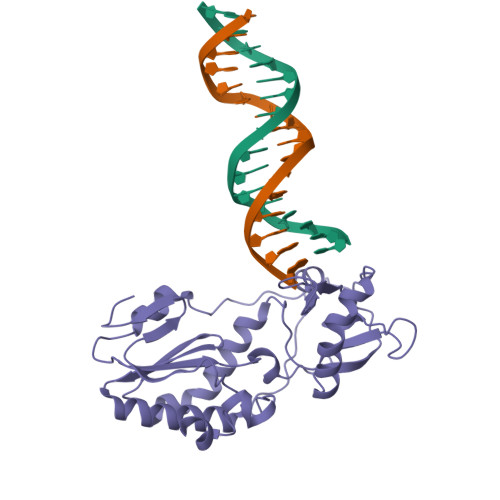
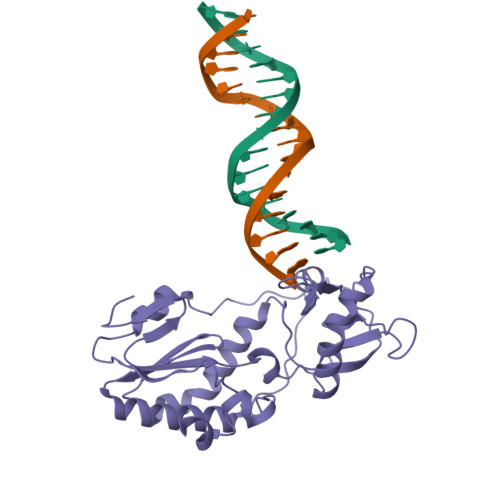
Staying Straight with A-tracts: A DNA Analog of the HIV-1 Polypurine Tract
Cote, M.L., Pflomm, M., Georgiadis, M.M.(2003) J Mol Biology 330: 57-74
- PubMed: 12818202
- DOI: https://doi.org/10.1016/s0022-2836(03)00554-0
- Primary Citation of Related Structures:
1N4L - PubMed Abstract:
The polypurine tract (PPT) from the HIV-1 genome is resistant to digestion by reverse transcriptase following (-)-strand synthesis and is used to prime (+)-strand synthesis during retroviral replication. We have determined the crystal structure of the asymmetric DNA/DNA analog16-mer duplex (CTTTTTAAAAGAAAAG/CTTTTCTTTTAAAAAG) comprising most of the "visible" portion of the RNA:DNA hybrid from the polypurine tract of HIV-1, which was previously reported in a complex with HIV-1 reverse transcriptase. Our 16-mer completely encompasses a 10-mer DNA duplex analog of the HIV-1 PPT. We report here a detailed analysis of our B' form 16-mer DNA structure, including three full pure A-tracts, as well as a comparative structural analysis with polypurine tract and other A-tract-containing nucleic acid structures. Our analysis reveals that the polypurine tract structures share structural features despite being different nucleic acid forms (i.e. DNA:DNA versus RNA:DNA). In addition, the previously reported A-tract-containing DNA molecules bound to topoisomerase I are remarkably similar to our polypurine tract 16-mer structure. On the basis of our analysis, we suggest that the specific topology of long pure A-tracts is remarkably comparable across a wide array of biological environments.
Organizational Affiliation:
Waksman Institute, Rutgers University, Piscataway, NJ 08854-8020, USA.