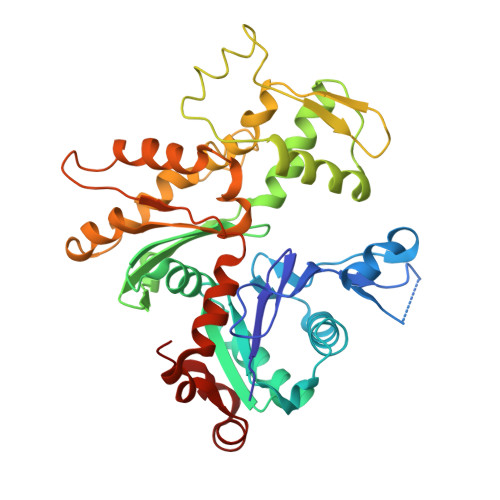
Polylysine induces an antiparallel actin dimer that nucleates filament assembly: crystal structure at 3.5-A resolution.
Bubb, M.R., Govindasamy, L., Yarmola, E.G., Vorobiev, S.M., Almo, S.C., Somasundaram, T., Chapman, M.S., Agbandje-Mckenna, M., Mckenna, R.(2002) J Biol Chem 277: 20999-21006
- PubMed: 11932258
- DOI: https://doi.org/10.1074/jbc.M201371200
- Primary Citation of Related Structures:
1IJJ, 1LCU - PubMed Abstract:
An antiparallel actin dimer has been proposed to be an intermediate species during actin filament nucleation. We now show that latrunculin A, a marine natural product that inhibits actin polymerization, arrests polylysine-induced nucleation at the level of an antiparallel dimer, resulting in its accumulation. These dimers, when composed of pyrene-labeled actin subunits, give rise to a fluorescent excimer, permitting detection during polymerization in vitro. We report the crystallographic structure of the polylysine-actin-latrunculin A complex at 3.5-A resolution. The non-crystallographic contact is consistent with a dimeric structure and confirms the antiparallel orientation of its subunits. The crystallographic contacts reveal that the mobile DNase I binding loop of one subunit of a symmetry-related antiparallel actin dimer is partially stabilized in the interface between the two subunits of a second antiparallel dimer. These results provide a potential explanation for the paradoxical nucleation of actin filaments that have exclusively parallel subunits by a dimer containing antiparallel subunits.
Organizational Affiliation:
Research Service, Malcom Randall Department of Veterans Affairs Medical Center and the University of Florida College of Medicine, Gainesville, Florida 32608, USA. bubbmr@medicine.ufl.edu