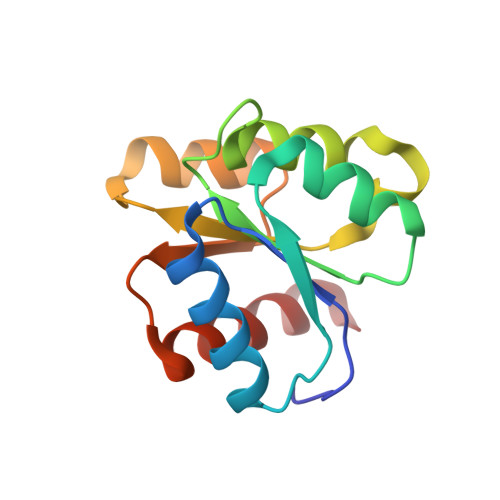
Three-dimensional structure of chemotactic Che Y protein in aqueous solution by nuclear magnetic resonance methods.
Santoro, J., Bruix, M., Pascual, J., Lopez, E., Serrano, L., Rico, M.(1995) J Mol Biol 247: 717-725
- PubMed: 7723026
- DOI: https://doi.org/10.1006/jmbi.1995.0175
- Primary Citation of Related Structures:
1CYE - PubMed Abstract:
The three-dimensional structure of chemotactic Che Y protein from Escherichia coli in aqueous solution has been determined by nuclear magnetic resonance (NMR) spectroscopy combined with restrained molecular dynamics calculations. A total of 20 converged structures were computed from 1545 conformationally relevant distance restraints derived from 1858 unambiguously assigned NOE cross-correlations. The resulting average pairwise root-mean-square deviation is 1.03 A for the backbone atoms and 1.69 A for all heavy atoms. If residues in the regions structurally least defined (1 to 5, 47 to 50, 76 to 79, 88 to 91 and 124 to 129) are excluded from the analysis, the root-mean-square deviations are reduced to 0.53 A and 1.23 A, respectively. The solution structure is closely similar to the refined X-ray crystal structure, except in the regions found to be less defined by NMR spectroscopy. The root-mean-square deviation between the average solution structure and the X-ray crystal structure is 0.92 A for the backbone residues (2 to 129). The highly refined solution structure determined herewith provides an essential background to delineate functionally important conformational changes brought about by different effectors.
Organizational Affiliation:
Instituto de Estructura de la Materia, CSIC, Madrid, Spain.